API reference
earthcarekit
earthcarekit
A Python package to simplify working with EarthCARE satellite data
See also:
Copyright © 2025 TROPOS
Cmap
Bases: ListedColormap
Colormap with categorical, gradient, and circular support.
This subclass of matplotlib.colors.ListedColormap adds utilities for
continuous and categorical color mappings. Supports labels, ticks,
normalization, blending, and transparency adjustments.
Attributes:
| Name | Type | Description |
|---|---|---|
categorical |
bool
|
Whether the colormap is discrete/categorical. |
gradient |
bool
|
Whether the colormap was generated from a gradient. |
circular |
bool
|
Whether the colormap wraps around cyclically. |
ticks |
list[float]
|
Optional tick positions for categorical plots. |
labels |
list[str]
|
Optional labels corresponding to ticks. |
norm |
Normalize | None
|
Normalization strategy for value mapping. |
values |
list
|
Associated values for categorical mapping. |
Source code in earthcarekit/plot/color/colormap/cmap.py
70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 | |
__init__
__init__(
colors,
name="colormap",
N=None,
categorical=False,
ticks=None,
labels=None,
norm=None,
values=None,
gradient=False,
circular=False,
)
Initialize a Cmap.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
colors
|
Sequence of colors (strings or ColorLike objects). |
required | |
name
|
str
|
Name of the colormap. Defaults to "colormap". |
'colormap'
|
N
|
int | None
|
Number of discrete colors. Defaults to None. |
None
|
categorical
|
bool
|
Whether the colormap is discrete/categorical. Defaults to False. |
False
|
ticks
|
list[float] | None
|
Optional tick positions for categorical plots. Defaults to None. |
None
|
labels
|
list[str] | None
|
Optional labels corresponding to ticks. Defaults to None. |
None
|
norm
|
Normalize | None
|
Normalization strategy for value mapping. Defaults to None. |
None
|
gradient
|
bool
|
If True, generate intermediate gradient colors. Defaults to False. |
False
|
circular
|
bool
|
If True, colormap wraps around cyclically. Defaults to False. |
False
|
Source code in earthcarekit/plot/color/colormap/cmap.py
__new__
Allow instantiation from an existing Colormap or standard arguments.
blend
Return a copy of the colormap blended with a second color.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
value
|
float
|
Blend factor in the range [0, 1]. |
required |
blend_color
|
Color | str
|
Color to blend with. |
'white'
|
Returns:
| Name | Type | Description |
|---|---|---|
Cmap |
Cmap
|
Blended colormap. |
Source code in earthcarekit/plot/color/colormap/cmap.py
from_colormap
classmethod
Create a Cmap instance from an existing Matplotlib colormap.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
cmap
|
Colormap
|
Source colormap to convert. |
required |
N
|
int
|
Number of discrete colors (if needed, e.g, for categorical colormaps with limited number of colors). Defaults to 256. |
256
|
Returns:
| Name | Type | Description |
|---|---|---|
Cmap |
Cmap
|
New colormap. |
Source code in earthcarekit/plot/color/colormap/cmap.py
set_alpha
Return a copy of the colormap with modified alpha transparency.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
value
|
float
|
Alpha value in the range [0, 1]. |
required |
Returns:
| Name | Type | Description |
|---|---|---|
Cmap |
Cmap
|
Colormap with updated transparency. |
Source code in earthcarekit/plot/color/colormap/cmap.py
to_categorical
Convert a colormap to categorical.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
values_to_labels
|
dict | int
|
Mapping from values to labels, or number of categories if int. |
required |
endpoint
|
bool | None
|
Whether the last color is included at 1.0. |
None
|
use_discrete
|
bool | None
|
If True, use the colormap's defined colors directly rather than sampling across its range. |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
Cmap |
Cmap
|
Categorical version of the colormap. |
Source code in earthcarekit/plot/color/colormap/cmap.py
to_discrete
Convert a colormap to a discretized version of itself.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
n
|
int
|
Number of steps (i.e., discrete colors). |
required |
Returns:
| Name | Type | Description |
|---|---|---|
Cmap |
Cmap
|
Discretized version of the colormap. |
Source code in earthcarekit/plot/color/colormap/cmap.py
Color
dataclass
Bases: str
Represents a color with convenient conversion, blending, and alpha support.
Extends str to store a color as a hex string while providing methods
to access RGB/RGBA, set transparency, blend with other colors, and
normalize input from various formats.
Attributes:
| Name | Type | Description |
|---|---|---|
input |
ColorLike
|
Original input used to create the color. |
name |
str | None
|
Optional name of the color. |
is_normalized |
bool
|
Whether the color values are normalized (0-1). |
Source code in earthcarekit/plot/color/color.py
54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 | |
__hash__
__init__
Initialize Color attributes.
Source code in earthcarekit/plot/color/color.py
__new__
Create a Color instance from a color-like input.
Source code in earthcarekit/plot/color/color.py
blend
Returns the same color blended with a second color.
Source code in earthcarekit/plot/color/color.py
from_optional
classmethod
Parses optional color input and returns a Color instance or None.
Source code in earthcarekit/plot/color/color.py
get_best_bw_contrast_color
Return black or white color depending on best contrast according to WCAG 2.0.
See https://www.w3.org/TR/WCAG20/
is_close_to_white
set_alpha
Returns the same color with the given transparency alpha value applied.
Source code in earthcarekit/plot/color/color.py
CurtainFigure
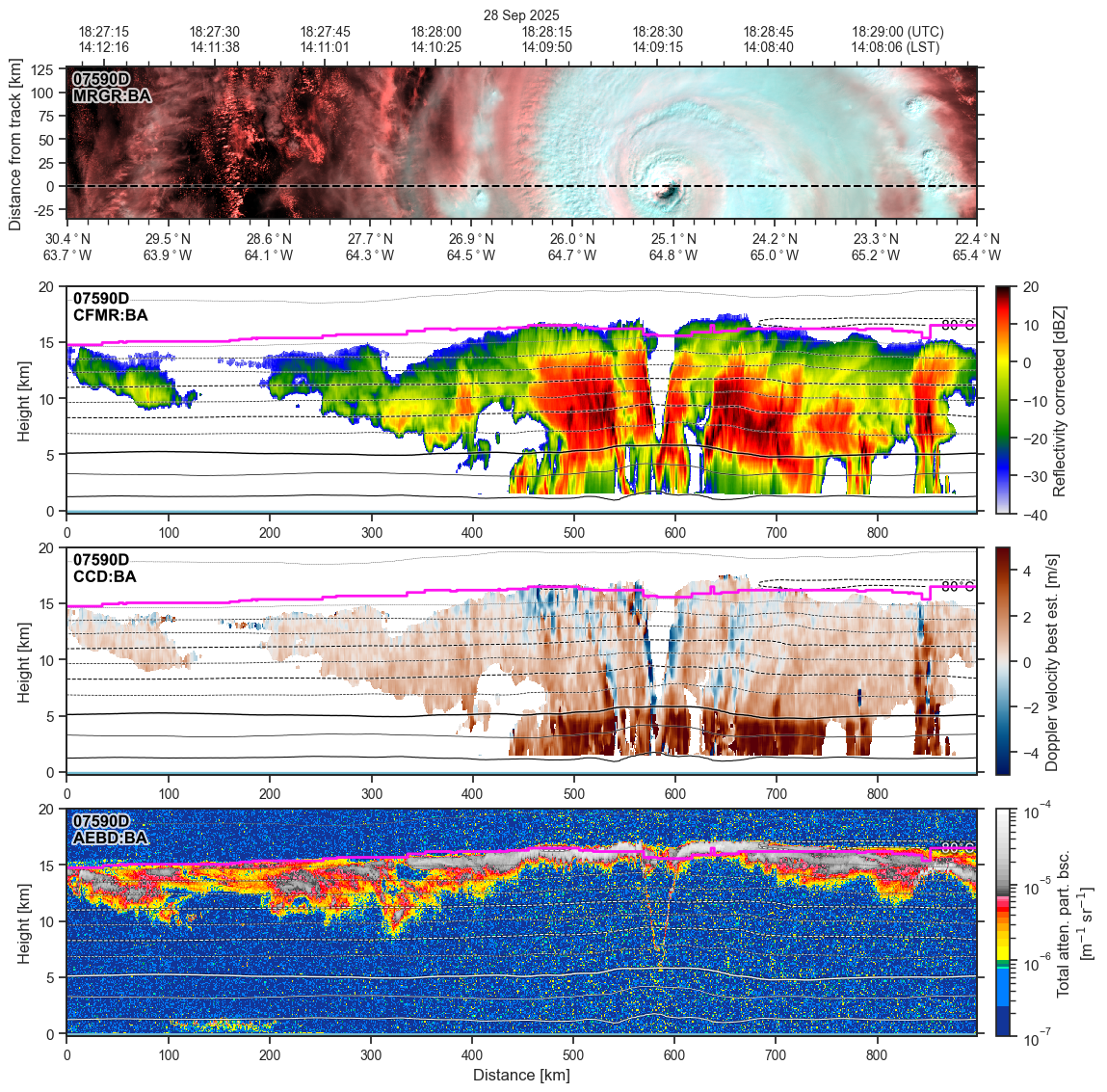
Figure object for displaying EarthCARE curtain data (e.g., ATLID and CPR L1/L2 profiles) along the satellite track.
This class sets up a horizontal-along-track or time vs. vertical-height plot (a "curtain" view), for profiling atmospheric quantities retrieved from ground-based or nadir-viewing air/space-bourne instruments (like EarthCARE). It displays dual top/bottom x-axes (e.g., geolocation and time), and left/right y-axes for height labels.
Attributes:
| Name | Type | Description |
|---|---|---|
ax |
Axes | None
|
Existing matplotlib axes to plot on; if not provided, a new figure and axes will be created. Defaults to None. |
figsize |
tuple[float, float]
|
Size of the figure in inches. Defaults to (FIGURE_WIDTH_CURTAIN, FIGURE_HEIGHT_CURTAIN). |
dpi |
int | None
|
Resolution of the figure in dots per inch. Defaults to None. |
title |
str | None
|
Title to display above the curtain plot. Defaults to None. |
ax_style_top |
AlongTrackAxisStyle | str
|
Style of the top x-axis, e.g., "geo", "time", or "frame". Defaults to "geo". |
ax_style_bottom |
AlongTrackAxisStyle | str
|
Style of the bottom x-axis, e.g., "geo", "time", or "frame". Defaults to "time". |
num_ticks |
int
|
Maximum number of tick marks to be place along the x-axis. Defaults to 10. |
show_height_left |
bool
|
Whether to show height labels on the left y-axis. Defaults to True. |
show_height_right |
bool
|
Whether to show height labels on the right y-axis. Defaults to False. |
mode |
Literal['exact', 'fast']
|
Curtain plotting mode. Use "fast" to speed up plotting by coarsening data to at least |
min_num_profiles |
int
|
Minimum number of profiles to keep when using "fast" mode. Defaults to 1000. |
Source code in earthcarekit/plot/figure/curtain.py
146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030 1031 1032 1033 1034 1035 1036 1037 1038 1039 1040 1041 1042 1043 1044 1045 1046 1047 1048 1049 1050 1051 1052 1053 1054 1055 1056 1057 1058 1059 1060 1061 1062 1063 1064 1065 1066 1067 1068 1069 1070 1071 1072 1073 1074 1075 1076 1077 1078 1079 1080 1081 1082 1083 1084 1085 1086 1087 1088 1089 1090 1091 1092 1093 1094 1095 1096 1097 1098 1099 1100 1101 1102 1103 1104 1105 1106 1107 1108 1109 1110 1111 1112 1113 1114 1115 1116 1117 1118 1119 1120 1121 1122 1123 1124 1125 1126 1127 1128 1129 1130 1131 1132 1133 1134 1135 1136 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 1148 1149 1150 1151 1152 1153 1154 1155 1156 1157 1158 1159 1160 1161 1162 1163 1164 1165 1166 1167 1168 1169 1170 1171 1172 1173 1174 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 1192 1193 1194 1195 1196 1197 1198 1199 1200 1201 1202 1203 1204 1205 1206 1207 1208 1209 1210 1211 1212 1213 1214 1215 1216 1217 1218 1219 1220 1221 1222 1223 1224 1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 1247 1248 1249 1250 1251 1252 1253 1254 1255 1256 1257 1258 1259 1260 1261 1262 1263 1264 1265 1266 1267 1268 1269 1270 1271 1272 1273 1274 1275 1276 1277 1278 1279 1280 1281 1282 1283 1284 1285 1286 1287 1288 1289 1290 1291 1292 1293 1294 1295 1296 1297 1298 1299 1300 1301 1302 1303 1304 1305 1306 1307 1308 1309 1310 1311 1312 1313 1314 1315 1316 1317 1318 1319 1320 1321 1322 1323 1324 1325 1326 1327 1328 1329 1330 1331 1332 1333 1334 1335 1336 1337 1338 1339 1340 1341 1342 1343 1344 1345 1346 1347 1348 1349 1350 1351 1352 1353 1354 1355 1356 1357 1358 1359 1360 1361 1362 1363 1364 1365 1366 1367 1368 1369 1370 1371 1372 1373 1374 1375 1376 1377 1378 1379 1380 1381 1382 1383 1384 1385 1386 1387 1388 1389 1390 1391 1392 1393 1394 1395 1396 1397 1398 1399 1400 1401 1402 1403 1404 1405 1406 1407 1408 1409 1410 1411 1412 1413 1414 1415 1416 1417 1418 1419 1420 1421 1422 1423 1424 1425 1426 1427 1428 1429 1430 1431 1432 1433 1434 1435 1436 1437 1438 1439 1440 1441 1442 1443 1444 1445 1446 1447 1448 1449 1450 1451 1452 1453 1454 1455 1456 1457 1458 1459 1460 1461 1462 1463 1464 1465 1466 1467 1468 1469 1470 1471 1472 1473 1474 1475 1476 1477 1478 1479 1480 1481 1482 1483 1484 1485 1486 1487 1488 1489 1490 1491 1492 1493 1494 1495 1496 1497 1498 1499 1500 1501 1502 1503 1504 1505 1506 1507 1508 1509 1510 1511 1512 1513 1514 1515 1516 1517 1518 1519 1520 1521 1522 1523 1524 1525 1526 1527 1528 1529 1530 1531 1532 1533 1534 1535 1536 1537 1538 1539 1540 1541 1542 1543 1544 1545 1546 1547 1548 1549 1550 1551 1552 1553 1554 1555 1556 1557 1558 1559 1560 1561 1562 1563 1564 1565 1566 1567 1568 1569 1570 1571 1572 1573 1574 1575 1576 1577 1578 1579 1580 1581 1582 1583 1584 1585 1586 1587 1588 1589 1590 1591 1592 1593 1594 1595 1596 1597 1598 1599 1600 1601 1602 1603 1604 1605 1606 1607 1608 1609 1610 1611 1612 1613 1614 1615 1616 1617 1618 1619 1620 1621 1622 1623 1624 1625 1626 1627 1628 1629 1630 1631 1632 1633 1634 1635 1636 1637 1638 1639 1640 1641 1642 1643 1644 1645 1646 1647 1648 1649 1650 1651 1652 1653 1654 1655 1656 1657 1658 1659 1660 1661 1662 1663 1664 1665 1666 1667 1668 1669 1670 1671 1672 1673 1674 1675 1676 1677 1678 1679 1680 1681 1682 1683 1684 1685 1686 1687 1688 1689 1690 1691 1692 1693 1694 1695 1696 1697 1698 1699 1700 1701 1702 1703 1704 1705 1706 1707 1708 1709 1710 1711 1712 1713 1714 1715 1716 1717 | |
ecplot
ecplot(
ds,
var,
*,
time_var=TIME_VAR,
height_var=HEIGHT_VAR,
lat_var=TRACK_LAT_VAR,
lon_var=TRACK_LON_VAR,
temperature_var=TEMP_CELSIUS_VAR,
along_track_dim=ALONG_TRACK_DIM,
values=None,
time=None,
height=None,
latitude=None,
longitude=None,
values_temperature=None,
site=None,
radius_km=100.0,
mark_closest_profile=False,
show_info=True,
show_info_orbit_and_frame=True,
show_info_file_type=True,
show_info_baseline=True,
info_text_orbit_and_frame=None,
info_text_file_type=None,
info_text_baseline=None,
show_radius=True,
info_text_loc=None,
value_range="default",
log_scale=None,
norm=None,
time_range=None,
height_range=(0, 40000.0),
label=None,
units=None,
cmap=None,
colorbar=True,
colorbar_ticks=None,
colorbar_tick_labels=None,
colorbar_position="right",
colorbar_alignment="center",
colorbar_width=DEFAULT_COLORBAR_WIDTH,
colorbar_spacing=0.2,
colorbar_length_ratio="100%",
colorbar_label_outside=True,
colorbar_ticks_outside=True,
colorbar_ticks_both=False,
rolling_mean=None,
selection_time_range=None,
selection_color=Color("ec:earthcare"),
selection_linestyle="dashed",
selection_linewidth=2.5,
selection_highlight=False,
selection_highlight_inverted=True,
selection_highlight_color=Color("white"),
selection_highlight_alpha=0.5,
selection_max_time_margin=None,
ax_style_top=None,
ax_style_bottom=None,
show_temperature=False,
mode=None,
min_num_profiles=5000,
mark_profiles_at=None,
mark_profiles_at_color=None,
mark_profiles_at_linestyle="solid",
mark_profiles_at_linewidth=2.5,
label_length=40,
**kwargs
)
Plot a vertical curtain (i.e. cross-section) of a variable along the satellite track a EarthCARE dataset.
This method collections all required data from a EarthCARE xarray.dataset, such as time, height, latitude and longitude.
It supports various forms of customization through the use of arguments listed below.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
Dataset
|
The EarthCARE dataset from with data will be plotted. |
required |
var
|
str
|
Name of the variable to plot. |
required |
time_var
|
str
|
Name of the time variable. Defaults to TIME_VAR. |
TIME_VAR
|
height_var
|
str
|
Name of the height variable. Defaults to HEIGHT_VAR. |
HEIGHT_VAR
|
lat_var
|
str
|
Name of the latitude variable. Defaults to TRACK_LAT_VAR. |
TRACK_LAT_VAR
|
lon_var
|
str
|
Name of the longitude variable. Defaults to TRACK_LON_VAR. |
TRACK_LON_VAR
|
temperature_var
|
str
|
Name of the temperature variable; ignored if |
TEMP_CELSIUS_VAR
|
along_track_dim
|
str
|
Dimension name representing the along-track direction. Defaults to ALONG_TRACK_DIM. |
ALONG_TRACK_DIM
|
values
|
NDArray | None
|
Data values to be used instead of values found in the |
None
|
time
|
NDArray | None
|
Time values to be used instead of values found in the |
None
|
height
|
NDArray | None
|
Height values to be used instead of values found in the |
None
|
latitude
|
NDArray | None
|
Latitude values to be used instead of values found in the |
None
|
longitude
|
NDArray | None
|
Longitude values to be used instead of values found in the |
None
|
values_temperature
|
NDArray | None
|
Temperature values to be used instead of values found in the |
None
|
site
|
str | GroundSite | None
|
Highlights data within |
None
|
radius_km
|
float
|
Radius around the ground site to highlight data from; ignored if |
100.0
|
mark_closest_profile
|
bool
|
Mark the closest profile to the ground site in the plot; ignored if |
False
|
show_info
|
bool
|
If True, show text on the plot containing EarthCARE frame and baseline info. Defaults to True. |
True
|
info_text_loc
|
str | None
|
Place info text at a specific location of the plot, e.g. "upper right" or "lower left". Defaults to None. |
None
|
value_range
|
ValueRangeLike | None
|
Min and max range for the variable values. Defaults to None. |
'default'
|
log_scale
|
bool | None
|
Whether to apply a logarithmic color scale. Defaults to None. |
None
|
norm
|
Normalize | None
|
Matplotlib norm to use for color scaling. Defaults to None. |
None
|
time_range
|
TimeRangeLike | None
|
Time range to restrict the data for plotting. Defaults to None. |
None
|
height_range
|
DistanceRangeLike | None
|
Height range to restrict the data for plotting. Defaults to (0, 40e3). |
(0, 40000.0)
|
label
|
str | None
|
Label to use for colorbar. Defaults to None. |
None
|
units
|
str | None
|
Units of the variable to show in the colorbar label. Defaults to None. |
None
|
cmap
|
str | Colormap | None
|
Colormap to use for plotting. Defaults to None. |
None
|
colorbar
|
bool
|
Whether to display a colorbar. Defaults to True. |
True
|
colorbar_ticks
|
ArrayLike | None
|
Custom tick values for the colorbar. Defaults to None. |
None
|
colorbar_tick_labels
|
ArrayLike | None
|
Custom labels for the colorbar ticks. Defaults to None. |
None
|
rolling_mean
|
int | None
|
Apply rolling mean along time axis with this window size. Defaults to None. |
None
|
selection_time_range
|
TimeRangeLike | None
|
Time range to highlight as a selection; ignored if |
None
|
selection_color
|
_type_
|
Color for the selection range marker lines. Defaults to Color("ec:earthcare"). |
Color('ec:earthcare')
|
selection_linestyle
|
str | None
|
Line style for selection range markers. Defaults to "dashed". |
'dashed'
|
selection_linewidth
|
float | int | None
|
Line width for selection range markers. Defaults to 2.5. |
2.5
|
selection_highlight
|
bool
|
Whether to highlight the selection region by shading outside or inside areas. Defaults to False. |
False
|
selection_highlight_inverted
|
bool
|
If True and |
True
|
selection_highlight_color
|
str | None
|
If True and |
Color('white')
|
selection_highlight_alpha
|
float
|
If True and |
0.5
|
selection_max_time_margin
|
TimedeltaLike | Sequence[TimedeltaLike]
|
Zooms the time axis to a given maximum time from a selected time area. Defaults to None. |
None
|
ax_style_top
|
AlongTrackAxisStyle | str | None
|
Style for the top axis (e.g., geo, lat, lon, distance, time, utc, lst, none). Defaults to None. |
None
|
ax_style_bottom
|
AlongTrackAxisStyle | str | None
|
Style for the bottom axis (e.g., geo, lat, lon, distance, time, utc, lst, none). Defaults to None. |
None
|
show_temperature
|
bool
|
Whether to overlay temperature as contours; requires either |
False
|
mode
|
Literal['exact', 'fast'] | None
|
Overwrites the curtain plotting mode. Use "fast" to speed up plotting by coarsening data to at least |
None
|
min_num_profiles
|
int
|
Overwrites the minimum number of profiles to keep when using "fast" mode. Defaults to 1000. |
5000
|
mark_profiles_at
|
Sequence[TimestampLike] | None
|
Timestamps at which to mark vertical profiles. Defaults to None. |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
CurtainFigure |
CurtainFigure
|
The figure object containing the curtain plot. |
Example
Source code in earthcarekit/plot/figure/curtain.py
722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 | |
ecplot_contour
ecplot_contour(
ds,
var,
time_var=TIME_VAR,
height_var=HEIGHT_VAR,
levels=None,
label_format=None,
label_levels=None,
linewidths=1.5,
linestyles="solid",
colors="black",
zorder=3,
)
Adds contour lines to the plot.
Source code in earthcarekit/plot/figure/curtain.py
ecplot_elevation
ecplot_elevation(
ds,
var=ELEVATION_VAR,
time_var=TIME_VAR,
land_flag_var=LAND_FLAG_VAR,
color="ec:land",
color_water="ec:water",
legend_label=None,
legend_label_water=None,
)
Adds filled elevation/surface area to the plot.
Source code in earthcarekit/plot/figure/curtain.py
ecplot_hatch
ecplot_hatch(
ds,
var,
value_range,
time_var=TIME_VAR,
height_var=HEIGHT_VAR,
hatch="/////",
linewidth=1,
linewidth_border=0,
color="black",
color_border=None,
zorder=2,
legend_label=None,
)
Adds hatched/filled areas to the plot.
Source code in earthcarekit/plot/figure/curtain.py
ecplot_hatch_attenuated
Adds hatched area where ATLID "simple_classification" shows "attenuated" (-1).
Source code in earthcarekit/plot/figure/curtain.py
ecplot_height
ecplot_height(
ds,
var,
time_var=TIME_VAR,
linewidth=1.5,
linestyle="none",
color="black",
zorder=2.1,
marker="s",
markersize=1,
show_info=True,
info_text_loc=None,
legend_label=None,
)
Adds height line to the plot.
Source code in earthcarekit/plot/figure/curtain.py
ecplot_pressure
ecplot_pressure(
ds,
var=PRESSURE_VAR,
time_var=TIME_VAR,
height_var=HEIGHT_VAR,
label_format="%d hPa",
**kwargs
)
Adds pressure contour lines to the plot.
Source code in earthcarekit/plot/figure/curtain.py
ecplot_temperature
ecplot_temperature(
ds,
var=TEMP_CELSIUS_VAR,
label_format="$%.0f^{\\circ}$C",
label_levels=[-80, -40, 0],
levels=[-80, -70, -60, -50, -40, -30, -20, -10, 0, 10, 20],
linewidths=[0.75, 0.25, 0.5, 0.5, 0.75, 0.5, 0.75, 0.5, 1.0, 0.5, 0.75],
linestyles=[
"dashed",
"dashed",
"dashed",
"dashed",
"dashed",
"dashed",
"dashed",
"dashed",
"solid",
"solid",
"solid",
],
colors="black",
**kwargs
)
Adds temperature contour lines to the plot.
Source code in earthcarekit/plot/figure/curtain.py
ecplot_tropopause
ecplot_tropopause(
ds,
var=TROPOPAUSE_VAR,
time_var=TIME_VAR,
color="ec:tropopause",
linewidth=2,
linestyle="solid",
legend_label=None,
)
Adds tropopause line to the plot.
Source code in earthcarekit/plot/figure/curtain.py
invert_xaxis
invert_yaxis
plot_contour
plot_contour(
values,
time,
height,
label_levels=None,
label_format=None,
levels=None,
linewidths=1.5,
linestyles="solid",
colors="black",
zorder=2,
)
Adds contour lines to the plot.
Source code in earthcarekit/plot/figure/curtain.py
1102 1103 1104 1105 1106 1107 1108 1109 1110 1111 1112 1113 1114 1115 1116 1117 1118 1119 1120 1121 1122 1123 1124 1125 1126 1127 1128 1129 1130 1131 1132 1133 1134 1135 1136 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 1148 1149 1150 1151 1152 1153 1154 1155 1156 1157 1158 1159 1160 1161 1162 1163 1164 1165 1166 1167 1168 1169 1170 1171 1172 1173 1174 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 1192 1193 | |
plot_hatch
plot_hatch(
values,
time,
height,
value_range,
hatch="/////",
linewidth=1,
linewidth_border=0,
color="black",
color_border=None,
zorder=2,
legend_label=None,
)
Adds hatched/filled areas to the plot.
Source code in earthcarekit/plot/figure/curtain.py
1195 1196 1197 1198 1199 1200 1201 1202 1203 1204 1205 1206 1207 1208 1209 1210 1211 1212 1213 1214 1215 1216 1217 1218 1219 1220 1221 1222 1223 1224 1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 1247 1248 1249 1250 1251 1252 1253 1254 1255 1256 | |
plot_height
plot_height(
height,
time,
linewidth=1.5,
linestyle="solid",
color=None,
alpha=1.0,
zorder=2,
marker=None,
markersize=None,
fill=False,
legend_label=None,
)
Adds height line to the plot.
Source code in earthcarekit/plot/figure/curtain.py
save
save(
filename="",
filepath=None,
ds=None,
ds_filepath=None,
dpi="figure",
orbit_and_frame=None,
utc_timestamp=None,
use_utc_creation_timestamp=False,
site_name=None,
hmax=None,
radius=None,
extra=None,
transparent_outside=False,
verbose=True,
print_prefix="",
create_dirs=False,
transparent_background=False,
resolution=None,
**kwargs
)
Save a figure as an image or vector graphic to a file and optionally format the file name in a structured way using EarthCARE metadata.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
figure
|
Figure | HasFigure
|
A figure object ( |
required |
filename
|
str
|
The base name of the file. Can be extended based on other metadata provided. Defaults to empty string. |
''
|
filepath
|
str | None
|
The path where the image is saved. Can be extended based on other metadata provided. Defaults to None. |
None
|
ds
|
Dataset | None
|
A EarthCARE dataset from which metadata will be taken. Defaults to None. |
None
|
ds_filepath
|
str | None
|
A path to a EarthCARE product from which metadata will be taken. Defaults to None. |
None
|
pad
|
float
|
Extra padding (i.e., empty space) around the image in inches. Defaults to 0.1. |
required |
dpi
|
float | figure
|
The resolution in dots per inch. If 'figure', use the figure's dpi value. Defaults to None. |
'figure'
|
orbit_and_frame
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
utc_timestamp
|
TimestampLike | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
use_utc_creation_timestamp
|
bool
|
Whether the time of image creation should be included in the file name. Defaults to False. |
False
|
site_name
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
hmax
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
radius
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
resolution
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
extra
|
str | None
|
A custom string to be included in the file name. Defaults to None. |
None
|
transparent_outside
|
bool
|
Whether the area outside figures should be transparent. Defaults to False. |
False
|
verbose
|
bool
|
Whether the progress of image creation should be printed to the console. Defaults to True. |
True
|
print_prefix
|
str
|
A prefix string to all console messages. Defaults to "". |
''
|
create_dirs
|
bool
|
Whether images should be saved in a folder structure based on provided metadata. Defaults to False. |
False
|
transparent_background
|
bool
|
Whether the background inside and outside of figures should be transparent. Defaults to False. |
False
|
**kwargs
|
dict[str, Any]
|
Keyword arguments passed to wrapped function call of |
{}
|
Source code in earthcarekit/plot/figure/curtain.py
1648 1649 1650 1651 1652 1653 1654 1655 1656 1657 1658 1659 1660 1661 1662 1663 1664 1665 1666 1667 1668 1669 1670 1671 1672 1673 1674 1675 1676 1677 1678 1679 1680 1681 1682 1683 1684 1685 1686 1687 1688 1689 1690 1691 1692 1693 1694 1695 1696 1697 1698 1699 1700 1701 1702 1703 1704 1705 1706 1707 1708 1709 1710 1711 1712 1713 1714 1715 1716 1717 | |
to_texture
Convert the figure to a texture by removing all axis ticks, labels, annotations, and text.
Source code in earthcarekit/plot/figure/curtain.py
FileAgency
Bases: FileInfoEnum
Source code in earthcarekit/utils/read/product/file_info/agency.py
from_input
classmethod
Infers the EarthCARE product agency (i.e. ESA or JAXA) from a given file or dataset.
Source code in earthcarekit/utils/read/product/file_info/agency.py
FileLatency
Bases: FileInfoEnum
Source code in earthcarekit/utils/read/product/file_info/latency.py
from_input
classmethod
Infers the EarthCARE product latency indicator (i.e. N for Near-real time, O for Offline, X for not applicable) from a given name, file or dataset.
Source code in earthcarekit/utils/read/product/file_info/latency.py
FileType
Bases: FileInfoEnum
Source code in earthcarekit/utils/read/product/file_info/type.py
73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 | |
from_input
classmethod
Infers the EarthCARE product type from a given file or dataset.
Source code in earthcarekit/utils/read/product/file_info/type.py
GroundSite
dataclass
Class representing a geographic site (or ground station) with associated metadata.
Attributes:
| Name | Type | Description |
|---|---|---|
latitude |
float
|
Latitude of the site in decimal degrees. |
longitude |
float
|
Longitude of the site in decimal degrees. |
name |
str
|
Short name or identifier of the site. |
long_name |
str
|
Full descriptive name of the site. |
aliases |
list[str]
|
Alternative names or identifiers for the site. |
altitude |
float
|
Altitude of the site in meters above sea level. |
cloudnet_name |
str | None
|
Identifier string used in CloudNet file names, or None if not applicable. |
Source code in earthcarekit/utils/ground_sites.py
aliases
class-attribute
instance-attribute
Alternative names or identifiers for the site.
altitude
class-attribute
instance-attribute
Altitude of the site in meters above sea level.
cloudnet_name
class-attribute
instance-attribute
Identifier string used in CloudNet file names, or None if not applicable.
LineFigure
TODO: documentation
Source code in earthcarekit/plot/figure/line.py
70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 | |
invert_xaxis
invert_yaxis
save
save(
filename="",
filepath=None,
ds=None,
ds_filepath=None,
dpi="figure",
orbit_and_frame=None,
utc_timestamp=None,
use_utc_creation_timestamp=False,
site_name=None,
hmax=None,
radius=None,
extra=None,
transparent_outside=False,
verbose=True,
print_prefix="",
create_dirs=False,
transparent_background=False,
resolution=None,
**kwargs
)
Save a figure as an image or vector graphic to a file and optionally format the file name in a structured way using EarthCARE metadata.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
figure
|
Figure | HasFigure
|
A figure object ( |
required |
filename
|
str
|
The base name of the file. Can be extended based on other metadata provided. Defaults to empty string. |
''
|
filepath
|
str | None
|
The path where the image is saved. Can be extended based on other metadata provided. Defaults to None. |
None
|
ds
|
Dataset | None
|
A EarthCARE dataset from which metadata will be taken. Defaults to None. |
None
|
ds_filepath
|
str | None
|
A path to a EarthCARE product from which metadata will be taken. Defaults to None. |
None
|
pad
|
float
|
Extra padding (i.e., empty space) around the image in inches. Defaults to 0.1. |
required |
dpi
|
float | figure
|
The resolution in dots per inch. If 'figure', use the figure's dpi value. Defaults to None. |
'figure'
|
orbit_and_frame
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
utc_timestamp
|
TimestampLike | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
use_utc_creation_timestamp
|
bool
|
Whether the time of image creation should be included in the file name. Defaults to False. |
False
|
site_name
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
hmax
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
radius
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
resolution
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
extra
|
str | None
|
A custom string to be included in the file name. Defaults to None. |
None
|
transparent_outside
|
bool
|
Whether the area outside figures should be transparent. Defaults to False. |
False
|
verbose
|
bool
|
Whether the progress of image creation should be printed to the console. Defaults to True. |
True
|
print_prefix
|
str
|
A prefix string to all console messages. Defaults to "". |
''
|
create_dirs
|
bool
|
Whether images should be saved in a folder structure based on provided metadata. Defaults to False. |
False
|
transparent_background
|
bool
|
Whether the background inside and outside of figures should be transparent. Defaults to False. |
False
|
**kwargs
|
dict[str, Any]
|
Keyword arguments passed to wrapped function call of |
{}
|
Source code in earthcarekit/plot/figure/line.py
MapFigure
Figure object for displaying EarthCARE satellite track and/or imager swaths on a global map.
This class sets up a georeferenced map canvas using a range of cartographic projections and visual styles. It serves as the basis for plotting 2D swath data (e.g., from MSI) or simple satellite tracks, optionally with info labels, backgrounds, and other styling options.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ax
|
Axes | None
|
Existing matplotlib axes to plot on; if not provided, new axes will be created. Defaults to None. |
None
|
figsize
|
tuple[float, float]
|
Figure size in inches. Defaults to (FIGURE_MAP_WIDTH, FIGURE_MAP_HEIGHT). |
(FIGURE_MAP_WIDTH, FIGURE_MAP_HEIGHT)
|
dpi
|
int | None
|
Resolution of the figure in dots per inch. Defaults to None. |
None
|
title
|
str | None
|
Title to display on the map. Defaults to None. |
None
|
style
|
str | Literal['none', 'stock_img', 'gray', 'osm', 'satellite', 'mtg', 'msg', 'blue_marble', 'land_ocean', 'land_ocean_lakes_rivers']
|
Style of the map's background image. Defaults to "gray". |
'gray'
|
projection
|
str | Projection
|
Map projection to use; options include "platecarree", "perspective", "orthographic", or a custom |
Orthographic()
|
central_latitude
|
float | None
|
Latitude at the center of the projection. Defaults to None. |
None
|
central_longitude
|
float | None
|
Longitude at the center of the projection. Defaults to None. |
0.0
|
grid_color
|
ColorLike | None
|
Color of grid lines. Defaults to None. |
None
|
border_color
|
ColorLike | None
|
Color of border box around the map. Defaults to None. |
None
|
coastline_color
|
ColorLike | None
|
Color of coastlines. Defaults to None. |
None
|
show_grid
|
bool
|
Whether to show latitude/longitude grid lines. Defaults to True. |
True
|
show_top_labels
|
bool
|
Whether to show tick labels on the top axis. Defaults to True. |
True
|
show_bottom_labels
|
bool
|
Whether to show tick labels on the bottom axis. Defaults to True. |
True
|
show_right_labels
|
bool
|
Whether to show tick labels on the right axis. Defaults to True. |
True
|
show_left_labels
|
bool
|
Whether to show tick labels on the left axis. Defaults to True. |
True
|
show_text_time
|
bool
|
Whether to display a datetime info text above the plot. Defaults to True. |
True
|
show_text_frame
|
bool
|
Whether to display a EarthCARE frame info text above the plot. Defaults to True. |
True
|
show_text_overpass
|
bool
|
Whether to display ground site overpass info in the plot. Defaults to True. |
True
|
show_night_shade
|
bool
|
Whether to overlay the nighttime shading based on |
True
|
timestamp
|
TimestampLike | None
|
Time reference used for nightshade overlay. Defaults to None. |
None
|
extent
|
Iterable | None
|
Map extent given as [lon_min, lon_max, lat_min, lat_max]; overrides auto zoom. Defaults to None. |
None
|
lod
|
int | None
|
Level of detail for choosen background map style image; higher values increase complexity. Defaults to None (meaning automatic selection). |
None
|
coastlines_resolution
|
str
|
Resolution of coastlines to display; options are "10m", "50m", or "110m". Defaults to "110m". |
'110m'
|
azimuth
|
float
|
Rotation of the |
0
|
pad
|
float | list[float]
|
Padding applied when selecting a map extent. Defaults to 0.05. |
0.05
|
background_alpha
|
float
|
Transparency level of the background map style. Defaults to 1.0. |
1.0
|
Source code in earthcarekit/plot/figure/map.py
339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030 1031 1032 1033 1034 1035 1036 1037 1038 1039 1040 1041 1042 1043 1044 1045 1046 1047 1048 1049 1050 1051 1052 1053 1054 1055 1056 1057 1058 1059 1060 1061 1062 1063 1064 1065 1066 1067 1068 1069 1070 1071 1072 1073 1074 1075 1076 1077 1078 1079 1080 1081 1082 1083 1084 1085 1086 1087 1088 1089 1090 1091 1092 1093 1094 1095 1096 1097 1098 1099 1100 1101 1102 1103 1104 1105 1106 1107 1108 1109 1110 1111 1112 1113 1114 1115 1116 1117 1118 1119 1120 1121 1122 1123 1124 1125 1126 1127 1128 1129 1130 1131 1132 1133 1134 1135 1136 1137 1138 1139 1140 1141 1142 1143 1144 1145 1146 1147 1148 1149 1150 1151 1152 1153 1154 1155 1156 1157 1158 1159 1160 1161 1162 1163 1164 1165 1166 1167 1168 1169 1170 1171 1172 1173 1174 1175 1176 1177 1178 1179 1180 1181 1182 1183 1184 1185 1186 1187 1188 1189 1190 1191 1192 1193 1194 1195 1196 1197 1198 1199 1200 1201 1202 1203 1204 1205 1206 1207 1208 1209 1210 1211 1212 1213 1214 1215 1216 1217 1218 1219 1220 1221 1222 1223 1224 1225 1226 1227 1228 1229 1230 1231 1232 1233 1234 1235 1236 1237 1238 1239 1240 1241 1242 1243 1244 1245 1246 1247 1248 1249 1250 1251 1252 1253 1254 1255 1256 1257 1258 1259 1260 1261 1262 1263 1264 1265 1266 1267 1268 1269 1270 1271 1272 1273 1274 1275 1276 1277 1278 1279 1280 1281 1282 1283 1284 1285 1286 1287 1288 1289 1290 1291 1292 1293 1294 1295 1296 1297 1298 1299 1300 1301 1302 1303 1304 1305 1306 1307 1308 1309 1310 1311 1312 1313 1314 1315 1316 1317 1318 1319 1320 1321 1322 1323 1324 1325 1326 1327 1328 1329 1330 1331 1332 1333 1334 1335 1336 1337 1338 1339 1340 1341 1342 1343 1344 1345 1346 1347 1348 1349 1350 1351 1352 1353 1354 1355 1356 1357 1358 1359 1360 1361 1362 1363 1364 1365 1366 1367 1368 1369 1370 1371 1372 1373 1374 1375 1376 1377 1378 1379 1380 1381 1382 1383 1384 1385 1386 1387 1388 1389 1390 1391 1392 1393 1394 1395 1396 1397 1398 1399 1400 1401 1402 1403 1404 1405 1406 1407 1408 1409 1410 1411 1412 1413 1414 1415 1416 1417 1418 1419 1420 1421 1422 1423 1424 1425 1426 1427 1428 1429 1430 1431 1432 1433 1434 1435 1436 1437 1438 1439 1440 1441 1442 1443 1444 1445 1446 1447 1448 1449 1450 1451 1452 1453 1454 1455 1456 1457 1458 1459 1460 1461 1462 1463 1464 1465 1466 1467 1468 1469 1470 1471 1472 1473 1474 1475 1476 1477 1478 1479 1480 1481 1482 1483 1484 1485 1486 1487 1488 1489 1490 1491 1492 1493 1494 1495 1496 1497 1498 1499 1500 1501 1502 1503 1504 1505 1506 1507 1508 1509 1510 1511 1512 1513 1514 1515 1516 1517 1518 1519 1520 1521 1522 1523 1524 1525 1526 1527 1528 1529 1530 1531 1532 1533 1534 1535 1536 1537 1538 1539 1540 1541 1542 1543 1544 1545 1546 1547 1548 1549 1550 1551 1552 1553 1554 1555 1556 1557 1558 1559 1560 1561 1562 1563 1564 1565 1566 1567 1568 1569 1570 1571 1572 1573 1574 1575 1576 1577 1578 1579 1580 1581 1582 1583 1584 1585 1586 1587 1588 1589 1590 1591 1592 1593 1594 1595 1596 1597 1598 1599 1600 1601 1602 1603 1604 1605 1606 1607 1608 1609 1610 1611 1612 1613 1614 1615 1616 1617 1618 1619 1620 1621 1622 1623 1624 1625 1626 1627 1628 1629 1630 1631 1632 1633 1634 1635 1636 1637 1638 1639 1640 1641 1642 1643 1644 1645 1646 1647 1648 1649 1650 1651 1652 1653 1654 1655 1656 1657 1658 1659 1660 1661 1662 1663 1664 1665 1666 1667 1668 1669 1670 1671 1672 1673 1674 1675 1676 1677 1678 1679 1680 1681 1682 1683 1684 1685 1686 1687 1688 1689 1690 1691 1692 1693 1694 1695 1696 1697 1698 1699 1700 1701 1702 1703 1704 1705 1706 1707 1708 1709 1710 1711 1712 1713 1714 1715 1716 1717 1718 1719 1720 1721 1722 1723 1724 1725 1726 1727 1728 1729 1730 1731 1732 1733 1734 1735 1736 1737 1738 1739 1740 1741 1742 1743 1744 1745 1746 1747 1748 1749 1750 1751 1752 1753 1754 1755 1756 1757 1758 1759 1760 1761 1762 1763 1764 1765 1766 1767 1768 1769 1770 1771 1772 1773 1774 1775 1776 1777 1778 1779 1780 1781 1782 1783 1784 1785 1786 1787 1788 1789 1790 1791 1792 1793 1794 1795 1796 1797 1798 1799 1800 1801 1802 1803 1804 1805 1806 1807 1808 1809 1810 1811 1812 1813 1814 1815 1816 1817 1818 1819 1820 1821 1822 1823 1824 1825 1826 1827 1828 1829 1830 1831 1832 1833 1834 1835 1836 1837 1838 1839 1840 1841 1842 1843 1844 1845 1846 1847 1848 1849 1850 1851 1852 1853 1854 1855 1856 1857 1858 1859 1860 1861 1862 1863 1864 1865 1866 1867 1868 1869 1870 1871 1872 1873 1874 1875 1876 1877 1878 1879 1880 1881 1882 1883 1884 1885 1886 1887 1888 1889 1890 1891 1892 1893 1894 1895 1896 1897 1898 1899 1900 1901 1902 1903 1904 1905 1906 1907 1908 1909 1910 1911 1912 1913 1914 1915 1916 1917 1918 1919 1920 1921 1922 1923 1924 1925 1926 1927 1928 1929 1930 1931 1932 1933 1934 1935 1936 1937 1938 1939 1940 1941 1942 1943 1944 1945 1946 1947 1948 1949 1950 1951 1952 1953 1954 1955 1956 1957 1958 1959 1960 1961 1962 1963 1964 1965 1966 1967 1968 1969 1970 1971 1972 1973 1974 1975 1976 1977 1978 1979 1980 1981 1982 1983 1984 1985 1986 1987 1988 1989 1990 1991 1992 1993 1994 1995 1996 1997 1998 1999 2000 2001 2002 2003 2004 2005 2006 2007 2008 2009 2010 2011 2012 2013 2014 2015 2016 2017 2018 2019 2020 2021 2022 2023 2024 2025 2026 2027 2028 2029 2030 2031 2032 2033 2034 2035 2036 2037 2038 2039 2040 2041 2042 2043 2044 2045 2046 2047 2048 2049 2050 2051 2052 2053 2054 2055 2056 2057 2058 2059 2060 2061 2062 2063 2064 2065 2066 2067 2068 2069 2070 2071 2072 2073 2074 2075 2076 2077 2078 2079 2080 2081 2082 2083 2084 2085 2086 2087 2088 2089 2090 2091 2092 2093 2094 2095 2096 2097 2098 2099 2100 2101 2102 2103 2104 2105 2106 2107 2108 2109 2110 2111 2112 2113 2114 2115 2116 2117 2118 2119 2120 2121 2122 2123 2124 2125 2126 2127 2128 2129 2130 2131 2132 2133 2134 2135 2136 2137 2138 2139 2140 2141 2142 2143 2144 2145 2146 2147 2148 2149 2150 2151 2152 2153 2154 2155 2156 2157 2158 2159 2160 2161 2162 2163 2164 2165 2166 2167 2168 | |
ecplot
ecplot(
ds,
var=None,
*,
lat_var=TRACK_LAT_VAR,
lon_var=TRACK_LON_VAR,
swath_lat_var=SWATH_LAT_VAR,
swath_lon_var=SWATH_LON_VAR,
time_var=TIME_VAR,
along_track_dim=ALONG_TRACK_DIM,
across_track_dim=ACROSS_TRACK_DIM,
site=None,
radius_km=100.0,
time_range=None,
view="global",
zoom_tmin=None,
zoom_tmax=None,
color="ec:earthcare",
linewidth=3,
linestyle="solid",
color2="ec:blue",
linewidth2=None,
linestyle2=None,
cmap=None,
zoom_radius_km=None,
extent=None,
central_latitude=None,
central_longitude=None,
value_range="default",
log_scale=None,
norm=None,
colorbar=True,
pad=None,
show_text_time=None,
show_text_frame=None,
show_text_overpass=None,
colorbar_position="bottom",
colorbar_alignment="center",
colorbar_width=DEFAULT_COLORBAR_WIDTH,
colorbar_spacing=0.3,
colorbar_length_ratio="100%",
colorbar_label_outside=True,
colorbar_ticks_outside=True,
colorbar_ticks_both=False,
selection_max_time_margin=None
)
Plot the EarthCARE satellite track on a map, optionally showing a 2D swath variable if var is provided.
This method collects all required data from an EarthCARE xarray.Dataset.
If var is given, the corresponding swath variable is plotted on the map using a
color scale. Otherwise, the satellite ground track is plotted as a colored line.
If time_range or site is given, the selected track section within the selected time range or in proximity to ground sites are highlighted.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
Dataset
|
The EarthCARE dataset from which data will be plotted. |
required |
var
|
str | None
|
Name of a 2D swath variable to plot. If None, only the satellite ground track is shown. Defaults to None. |
None
|
lat_var
|
str
|
Name of the latitude variable for the along-track data. Defaults to TRACK_LAT_VAR. |
TRACK_LAT_VAR
|
lon_var
|
str
|
Name of the longitude variable for the along-track data. Defaults to TRACK_LON_VAR. |
TRACK_LON_VAR
|
swath_lat_var
|
str
|
Name of the latitude variable for the swath. Defaults to SWATH_LAT_VAR. |
SWATH_LAT_VAR
|
swath_lon_var
|
str
|
Name of the longitude variable for the swath. Defaults to SWATH_LON_VAR. |
SWATH_LON_VAR
|
time_var
|
str
|
Name of the time variable. Defaults to TIME_VAR. |
TIME_VAR
|
along_track_dim
|
str
|
Dimension name representing the along-track direction. Defaults to ALONG_TRACK_DIM. |
ALONG_TRACK_DIM
|
across_track_dim
|
str
|
Dimension name representing the across-track direction. Defaults to ACROSS_TRACK_DIM. |
ACROSS_TRACK_DIM
|
site
|
str | GroundSite | None
|
Highlights data within |
None
|
radius_km
|
float
|
Radius around the ground site to highlight data from; ignored if |
100.0
|
time_range
|
TimeRangeLike | None
|
Time range to highlight as selection area; ignored if |
None
|
view
|
Literal['global', 'data', 'overpass']
|
Map extent mode: "global" for full world, "data" for tight bounds, or "overpass" to zoom around |
'global'
|
zoom_tmin
|
TimestampLike | None
|
Optional lower time bound used for zooming map around track. Defaults to None. |
None
|
zoom_tmax
|
TimestampLike | None
|
Optional upper time bound used for zooming map around track. Defaults to None. |
None
|
color
|
ColorLike | None
|
Color used for selected section of the track or entire track if no selection. Defaults to "ec:earthcare". |
'ec:earthcare'
|
linewidth
|
float
|
Line width for selected track section. Defaults to 3. |
3
|
linestyle
|
str | None
|
Line style for selected track section. Defaults to "solid". |
'solid'
|
color2
|
ColorLike | None
|
Color used for unselected sections of the track. Defaults to "ec:blue". |
'ec:blue'
|
linewidth2
|
float
|
Line width for unselected sections. Defaults to None. |
None
|
linestyle2
|
str | None
|
Line style for unselected sections. Defaults to None. |
None
|
cmap
|
str | Cmap | None
|
Colormap to use when plotting a swath variable. Defaults to None. |
None
|
zoom_radius_km
|
float | None
|
If set, overrides map extent derived from |
None
|
extent
|
list[float] | None
|
Map extent in the form [lon_min, lon_max, lat_min, lat_max]. If given, overrides map extent derived from |
None
|
central_latitude
|
float | None
|
Central latitude used for the map projection. Defaults to None. |
None
|
central_longitude
|
float | None
|
Central longitude used for the map projection. Defaults to None. |
None
|
value_range
|
ValueRangeLike | None
|
Min and max range for the variable values; ignored if |
'default'
|
log_scale
|
bool | None
|
Whether to apply a logarithmic color scale to the variable. Defaults to None. |
None
|
norm
|
Normalize | None
|
Matplotlib norm to use for color scaling. Defaults to None. |
None
|
colorbar
|
bool
|
Whether to display a colorbar for the variable. Defaults to True. |
True
|
pad
|
float | list[float] | None
|
Padding around the map extent; ignored if |
None
|
show_text_time
|
bool | None
|
Whether to display the UTC time start and end of the selected track. Defaults to None. |
None
|
show_text_frame
|
bool | None
|
Whether to display EarthCARE frame information. Defaults to None. |
None
|
show_text_overpass
|
bool | None
|
Whether to display overpass site name and related info. Defaults to None. |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
MapFigure |
MapFigure
|
The figure object containing the map with track or swath. |
Example
Source code in earthcarekit/plot/figure/map.py
1396 1397 1398 1399 1400 1401 1402 1403 1404 1405 1406 1407 1408 1409 1410 1411 1412 1413 1414 1415 1416 1417 1418 1419 1420 1421 1422 1423 1424 1425 1426 1427 1428 1429 1430 1431 1432 1433 1434 1435 1436 1437 1438 1439 1440 1441 1442 1443 1444 1445 1446 1447 1448 1449 1450 1451 1452 1453 1454 1455 1456 1457 1458 1459 1460 1461 1462 1463 1464 1465 1466 1467 1468 1469 1470 1471 1472 1473 1474 1475 1476 1477 1478 1479 1480 1481 1482 1483 1484 1485 1486 1487 1488 1489 1490 1491 1492 1493 1494 1495 1496 1497 1498 1499 1500 1501 1502 1503 1504 1505 1506 1507 1508 1509 1510 1511 1512 1513 1514 1515 1516 1517 1518 1519 1520 1521 1522 1523 1524 1525 1526 1527 1528 1529 1530 1531 1532 1533 1534 1535 1536 1537 1538 1539 1540 1541 1542 1543 1544 1545 1546 1547 1548 1549 1550 1551 1552 1553 1554 1555 1556 1557 1558 1559 1560 1561 1562 1563 1564 1565 1566 1567 1568 1569 1570 1571 1572 1573 1574 1575 1576 1577 1578 1579 1580 1581 1582 1583 1584 1585 1586 1587 1588 1589 1590 1591 1592 1593 1594 1595 1596 1597 1598 1599 1600 1601 1602 1603 1604 1605 1606 1607 1608 1609 1610 1611 1612 1613 1614 1615 1616 1617 1618 1619 1620 1621 1622 1623 1624 1625 1626 1627 1628 1629 1630 1631 1632 1633 1634 1635 1636 1637 1638 1639 1640 1641 1642 1643 1644 1645 1646 1647 1648 1649 1650 1651 1652 1653 1654 1655 1656 1657 1658 1659 1660 1661 1662 1663 1664 1665 1666 1667 1668 1669 1670 1671 1672 1673 1674 1675 1676 1677 1678 1679 1680 1681 1682 1683 1684 1685 1686 1687 1688 1689 1690 1691 1692 1693 1694 1695 1696 1697 1698 1699 1700 1701 1702 1703 1704 1705 1706 1707 1708 1709 1710 1711 1712 1713 1714 1715 1716 1717 1718 1719 1720 1721 1722 1723 1724 1725 1726 1727 1728 1729 1730 1731 1732 1733 1734 1735 1736 1737 1738 1739 1740 1741 1742 1743 1744 1745 1746 1747 1748 1749 1750 1751 1752 1753 1754 1755 1756 1757 1758 1759 1760 1761 1762 1763 1764 1765 1766 1767 1768 1769 1770 1771 1772 1773 1774 1775 1776 1777 1778 1779 1780 1781 1782 1783 1784 1785 1786 1787 1788 1789 1790 1791 1792 1793 1794 1795 1796 1797 1798 1799 1800 1801 1802 1803 1804 1805 1806 1807 1808 1809 1810 1811 1812 1813 1814 1815 1816 1817 1818 1819 1820 1821 1822 1823 1824 1825 1826 1827 1828 1829 1830 1831 1832 1833 | |
save
save(
filename="",
filepath=None,
ds=None,
ds_filepath=None,
dpi="figure",
orbit_and_frame=None,
utc_timestamp=None,
use_utc_creation_timestamp=False,
site_name=None,
hmax=None,
radius=None,
extra=None,
transparent_outside=False,
verbose=True,
print_prefix="",
create_dirs=False,
transparent_background=False,
resolution=None,
**kwargs
)
Save a figure as an image or vector graphic to a file and optionally format the file name in a structured way using EarthCARE metadata.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
figure
|
Figure | HasFigure
|
A figure object ( |
required |
filename
|
str
|
The base name of the file. Can be extended based on other metadata provided. Defaults to empty string. |
''
|
filepath
|
str | None
|
The path where the image is saved. Can be extended based on other metadata provided. Defaults to None. |
None
|
ds
|
Dataset | None
|
A EarthCARE dataset from which metadata will be taken. Defaults to None. |
None
|
ds_filepath
|
str | None
|
A path to a EarthCARE product from which metadata will be taken. Defaults to None. |
None
|
pad
|
float
|
Extra padding (i.e., empty space) around the image in inches. Defaults to 0.1. |
required |
dpi
|
float | figure
|
The resolution in dots per inch. If 'figure', use the figure's dpi value. Defaults to None. |
'figure'
|
orbit_and_frame
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
utc_timestamp
|
TimestampLike | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
use_utc_creation_timestamp
|
bool
|
Whether the time of image creation should be included in the file name. Defaults to False. |
False
|
site_name
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
hmax
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
radius
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
resolution
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
extra
|
str | None
|
A custom string to be included in the file name. Defaults to None. |
None
|
transparent_outside
|
bool
|
Whether the area outside figures should be transparent. Defaults to False. |
False
|
verbose
|
bool
|
Whether the progress of image creation should be printed to the console. Defaults to True. |
True
|
print_prefix
|
str
|
A prefix string to all console messages. Defaults to "". |
''
|
create_dirs
|
bool
|
Whether images should be saved in a folder structure based on provided metadata. Defaults to False. |
False
|
transparent_background
|
bool
|
Whether the background inside and outside of figures should be transparent. Defaults to False. |
False
|
**kwargs
|
dict[str, Any]
|
Keyword arguments passed to wrapped function call of |
{}
|
Source code in earthcarekit/plot/figure/map.py
2099 2100 2101 2102 2103 2104 2105 2106 2107 2108 2109 2110 2111 2112 2113 2114 2115 2116 2117 2118 2119 2120 2121 2122 2123 2124 2125 2126 2127 2128 2129 2130 2131 2132 2133 2134 2135 2136 2137 2138 2139 2140 2141 2142 2143 2144 2145 2146 2147 2148 2149 2150 2151 2152 2153 2154 2155 2156 2157 2158 2159 2160 2161 2162 2163 2164 2165 2166 2167 2168 | |
set_view
Fits the plot extent to the given latitude and longitude values.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
latitude
|
ArrayLike
|
Latitude values. |
required |
longitude
|
ArrayLike
|
Longitude values. |
required |
pad
|
float | Iterable | None
|
Padding or margins around the given lat/lon values.
The padding is applied relative to the min/max difference along the respective lat/lon extent,
e.g., |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
Axes |
MapFigure
|
description |
Source code in earthcarekit/plot/figure/map.py
to_texture
Convert the figure to a texture by removing all axis ticks, labels, annotations, and text.
Source code in earthcarekit/plot/figure/map.py
ProductInfo
dataclass
Class storing all info gathered from a EarthCARE product's file path.
Attributes:
| Name | Type | Description |
|---|---|---|
mission_id |
FileMissionID
|
Mission ID (ECA = EarthCARE). |
agency |
FileAgency
|
Agency that generated the file (E = ESA, J = JAXA). |
latency |
FileLatency
|
Latency indicator (X = not applicable, N = near real-time, O = offline). |
baseline |
str
|
Two-letter product/processor version string (e.g., "BA"). |
file_type |
FileType
|
Full product name (10 characters, e.g., "ATL_EBD_2A"). |
start_sensing_time |
Timestamp
|
Start-time of data collection (i.e., time of first available data in the product). |
start_processing_time |
Timestamp
|
Start-time of processing (i.e., time at which creation of the product started). |
orbit_number |
int
|
Number of the orbit. |
frame_id |
str
|
Single letter identifier between A and H, indication the orbit segment (A,B,H = night frames; D,E,F = day frames; C,G = polar day/night frames). |
orbit_and_frame |
str
|
Six-character string with leading zeros combining orbit number and frame ID. |
name |
str
|
Full name of the product without file extension. |
filepath |
str
|
Local file path or empty string if not available. |
hdr_filepath |
str
|
Local header file path or empty string if not available. |
Source code in earthcarekit/utils/read/product/file_info/product_info.py
to_dataframe
ProfileData
dataclass
Container for atmospheric profile data.
Stores profile values together with their time/height bins and,
optionally, their coordinates and metadata in a consistent structure,
making profiles easier to handle, compare and visualise.
The object supports NumPy-style indexing based on its values attribute
following the convention: profile[time_index, height_index].
Attributes:
| Name | Type | Description |
|---|---|---|
values |
NDArray
|
Profile data, either a single vertical profile or a time series of profiles (time x height). |
height |
NDArray
|
Height bin centers, ascending. Can be fixed or vary with time. |
time |
NDArray
|
Timestamps corresponding to each profile. |
latitude |
NDArray | None
|
Ground latitudes for the profiles, optional. |
longitude |
NDArray | None
|
Ground longitudes for the profiles, optional. |
color |
str | None
|
Color for plotting, optional. |
label |
str | None
|
Variable label for plot annotations, optional. |
units |
str | None
|
Units for plot annotations, optional. |
platform |
str | None
|
Name or type of measurement platform/instrument, optional. |
error |
NDArray | None
|
Associated uncertainties for the profile values, optional. |
Source code in earthcarekit/utils/profile_data/profile_data.py
158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030 1031 1032 1033 1034 1035 1036 1037 1038 1039 1040 1041 1042 1043 1044 1045 1046 1047 1048 1049 1050 1051 1052 1053 1054 1055 1056 1057 1058 1059 1060 1061 1062 1063 1064 1065 1066 1067 1068 1069 1070 1071 1072 1073 1074 1075 1076 1077 1078 1079 1080 1081 1082 1083 1084 1085 1086 1087 1088 1089 1090 1091 1092 1093 1094 1095 1096 1097 1098 1099 1100 1101 1102 1103 1104 1105 1106 1107 1108 1109 1110 1111 1112 1113 1114 1115 1116 1117 1118 1119 | |
coarsen_mean
Returns downsampled profile data.
Source code in earthcarekit/utils/profile_data/profile_data.py
layer_mean
Returns layer mean values.
Source code in earthcarekit/utils/profile_data/profile_data.py
mean
Returns mean profile.
Source code in earthcarekit/utils/profile_data/profile_data.py
rebin_along_track
Rebins profiles to new time bins.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
latitude_bin_centers
|
ArrayLike
|
Target time bin centers as a 1D array (shape represents temporal dimension) |
required |
Returns:
| Name | Type | Description |
|---|---|---|
rebinned_profiles |
VerticalProfiles
|
Profiles rebinned along the temporal dimension according to |
Source code in earthcarekit/utils/profile_data/profile_data.py
rebin_height
Rebins profiles to new height bins.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
new_height
|
ndarray
|
Target height bin centers as a 1D array (shape represents vertical dimension) |
required |
Returns:
| Name | Type | Description |
|---|---|---|
rebinned_profiles |
VerticalProfiles
|
Profiles rebinned along the vertical dimension according to |
Source code in earthcarekit/utils/profile_data/profile_data.py
rebin_time
Rebins profiles to new time bins.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
time_bin_centers
|
Iterable[TimestampLike] | ArrayLike
|
Target time bin centers as a 1D array (shape represents temporal dimension) |
required |
Returns:
| Name | Type | Description |
|---|---|---|
rebinned_profiles |
VerticalProfiles
|
Profiles rebinned along the temporal dimension according to |
Source code in earthcarekit/utils/profile_data/profile_data.py
rolling_mean
Returns mean profile.
Source code in earthcarekit/utils/profile_data/profile_data.py
select_height_range
Returns only data within the specified height_range.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
height_range
|
DistanceRangeLike
|
Pair of minimum and maximum height in meters. |
required |
pad_idx
|
int
|
Number of indexes that will be appended to the result before and after given height range. Defaults to 0. |
0
|
Returns:
| Name | Type | Description |
|---|---|---|
ProfileData |
ProfileData
|
New instance of ProfileData filtered by given height range. |
Source code in earthcarekit/utils/profile_data/profile_data.py
select_time_range
Returns only data within the specified time_range.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
time_range
|
TimeRangeLike | None
|
Pair of minimum and maximum timestamps or None. |
required |
pad_idx
|
int
|
Number of indexes that will be appended to the result before and after given time range. Defaults to 0. |
required |
Returns:
| Name | Type | Description |
|---|---|---|
ProfileData |
ProfileData
|
New instance of ProfileData filtered by given time range. |
Source code in earthcarekit/utils/profile_data/profile_data.py
std
Returns standard deviation profile.
Source code in earthcarekit/utils/profile_data/profile_data.py
ProfileFigure
Source code in earthcarekit/plot/figure/profile.py
120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 | |
invert_xaxis
invert_yaxis
plot
plot(
profiles=None,
*,
values=None,
time=None,
height=None,
latitude=None,
longitude=None,
error=None,
label=None,
units=None,
value_range=(0, None),
height_range=None,
time_range=None,
selection_height_range=None,
show_mean=True,
show_std=True,
show_min=False,
show_max=False,
show_sem=False,
show_error=False,
color=None,
alpha=1.0,
linestyle="solid",
linewidth=1.5,
ribbon_alpha=0.2,
show_grid=None,
zorder=1,
legend_label=None,
show_legend=None,
show_steps=DEFAULT_PROFILE_SHOW_STEPS
)
TODO: documentation
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
profiles
|
ProfileData | None
|
description. Defaults to None. |
None
|
values
|
NDArray | None
|
description. Defaults to None. |
None
|
time
|
NDArray | None
|
description. Defaults to None. |
None
|
height
|
NDArray | None
|
description. Defaults to None. |
None
|
latitude
|
NDArray | None
|
description. Defaults to None. |
None
|
longitude
|
NDArray | None
|
description. Defaults to None. |
None
|
error
|
NDArray | None
|
description. Defaults to None. |
None
|
label
|
str | None
|
description. Defaults to None. |
None
|
units
|
str | None
|
description. Defaults to None. |
None
|
value_range
|
ValueRangeLike | None
|
description. Defaults to (0, None). |
(0, None)
|
height_range
|
DistanceRangeLike | None
|
description. Defaults to None. |
None
|
time_range
|
TimeRangeLike | None
|
description. Defaults to None. |
None
|
selection_height_range
|
DistanceRangeLike | None
|
description. Defaults to None. |
None
|
show_mean
|
bool
|
description. Defaults to True. |
True
|
show_std
|
bool
|
description. Defaults to True. |
True
|
show_min
|
bool
|
description. Defaults to False. |
False
|
show_max
|
bool
|
description. Defaults to False. |
False
|
show_sem
|
bool
|
description. Defaults to False. |
False
|
show_error
|
bool
|
description. Defaults to False. |
False
|
color
|
str | ColorLike | None
|
description. Defaults to None. |
None
|
alpha
|
float
|
description. Defaults to 1.0. |
1.0
|
linestyle
|
str
|
description. Defaults to "solid". |
'solid'
|
linewidth
|
Number
|
description. Defaults to 1.5. |
1.5
|
ribbon_alpha
|
float
|
description. Defaults to 0.2. |
0.2
|
show_grid
|
bool | None
|
description. Defaults to None. |
None
|
zorder
|
Number | None
|
description. Defaults to 1. |
1
|
legend_label
|
str | None
|
description. Defaults to None. |
None
|
show_legend
|
bool | None
|
description. Defaults to None. |
None
|
show_steps
|
bool
|
description. Defaults to DEFAULT_PROFILE_SHOW_STEPS. |
DEFAULT_PROFILE_SHOW_STEPS
|
Raises:
| Type | Description |
|---|---|
ValueError
|
description |
ValueError
|
description |
Returns:
| Name | Type | Description |
|---|---|---|
ProfileFigure |
ProfileFigure
|
description |
Source code in earthcarekit/plot/figure/profile.py
269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 | |
save
save(
filename="",
filepath=None,
ds=None,
ds_filepath=None,
dpi="figure",
orbit_and_frame=None,
utc_timestamp=None,
use_utc_creation_timestamp=False,
site_name=None,
hmax=None,
radius=None,
extra=None,
transparent_outside=False,
verbose=True,
print_prefix="",
create_dirs=False,
transparent_background=False,
resolution=None,
**kwargs
)
Save a figure as an image or vector graphic to a file and optionally format the file name in a structured way using EarthCARE metadata.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
figure
|
Figure | HasFigure
|
A figure object ( |
required |
filename
|
str
|
The base name of the file. Can be extended based on other metadata provided. Defaults to empty string. |
''
|
filepath
|
str | None
|
The path where the image is saved. Can be extended based on other metadata provided. Defaults to None. |
None
|
ds
|
Dataset | None
|
A EarthCARE dataset from which metadata will be taken. Defaults to None. |
None
|
ds_filepath
|
str | None
|
A path to a EarthCARE product from which metadata will be taken. Defaults to None. |
None
|
pad
|
float
|
Extra padding (i.e., empty space) around the image in inches. Defaults to 0.1. |
required |
dpi
|
float | figure
|
The resolution in dots per inch. If 'figure', use the figure's dpi value. Defaults to None. |
'figure'
|
orbit_and_frame
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
utc_timestamp
|
TimestampLike | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
use_utc_creation_timestamp
|
bool
|
Whether the time of image creation should be included in the file name. Defaults to False. |
False
|
site_name
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
hmax
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
radius
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
resolution
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
extra
|
str | None
|
A custom string to be included in the file name. Defaults to None. |
None
|
transparent_outside
|
bool
|
Whether the area outside figures should be transparent. Defaults to False. |
False
|
verbose
|
bool
|
Whether the progress of image creation should be printed to the console. Defaults to True. |
True
|
print_prefix
|
str
|
A prefix string to all console messages. Defaults to "". |
''
|
create_dirs
|
bool
|
Whether images should be saved in a folder structure based on provided metadata. Defaults to False. |
False
|
transparent_background
|
bool
|
Whether the background inside and outside of figures should be transparent. Defaults to False. |
False
|
**kwargs
|
dict[str, Any]
|
Keyword arguments passed to wrapped function call of |
{}
|
Source code in earthcarekit/plot/figure/profile.py
SwathFigure
TODO: documentation
Source code in earthcarekit/plot/figure/swath.py
41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 516 517 518 519 520 521 522 523 524 525 526 527 528 529 530 531 532 533 534 535 536 537 538 539 540 541 542 543 544 545 546 547 548 549 550 551 552 553 554 555 556 557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 | |
invert_xaxis
invert_yaxis
plot_contour
plot_contour(
values,
time,
latitude,
longitude,
nadir_index,
label_levels=None,
label_format=None,
levels=None,
linewidths=1.5,
linestyles="solid",
colors="black",
zorder=2,
show_labels=True,
)
Adds contour lines to the plot.
Source code in earthcarekit/plot/figure/swath.py
420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 465 466 467 468 469 470 471 472 473 474 475 476 477 478 479 480 481 482 483 484 485 486 487 488 489 490 491 492 493 494 495 496 497 498 499 500 501 502 503 504 505 506 507 508 509 510 511 512 513 514 515 | |
save
save(
filename="",
filepath=None,
ds=None,
ds_filepath=None,
dpi="figure",
orbit_and_frame=None,
utc_timestamp=None,
use_utc_creation_timestamp=False,
site_name=None,
hmax=None,
radius=None,
extra=None,
transparent_outside=False,
verbose=True,
print_prefix="",
create_dirs=False,
transparent_background=False,
resolution=None,
**kwargs
)
Save a figure as an image or vector graphic to a file and optionally format the file name in a structured way using EarthCARE metadata.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
figure
|
Figure | HasFigure
|
A figure object ( |
required |
filename
|
str
|
The base name of the file. Can be extended based on other metadata provided. Defaults to empty string. |
''
|
filepath
|
str | None
|
The path where the image is saved. Can be extended based on other metadata provided. Defaults to None. |
None
|
ds
|
Dataset | None
|
A EarthCARE dataset from which metadata will be taken. Defaults to None. |
None
|
ds_filepath
|
str | None
|
A path to a EarthCARE product from which metadata will be taken. Defaults to None. |
None
|
pad
|
float
|
Extra padding (i.e., empty space) around the image in inches. Defaults to 0.1. |
required |
dpi
|
float | figure
|
The resolution in dots per inch. If 'figure', use the figure's dpi value. Defaults to None. |
'figure'
|
orbit_and_frame
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
utc_timestamp
|
TimestampLike | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
use_utc_creation_timestamp
|
bool
|
Whether the time of image creation should be included in the file name. Defaults to False. |
False
|
site_name
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
hmax
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
radius
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
resolution
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
extra
|
str | None
|
A custom string to be included in the file name. Defaults to None. |
None
|
transparent_outside
|
bool
|
Whether the area outside figures should be transparent. Defaults to False. |
False
|
verbose
|
bool
|
Whether the progress of image creation should be printed to the console. Defaults to True. |
True
|
print_prefix
|
str
|
A prefix string to all console messages. Defaults to "". |
''
|
create_dirs
|
bool
|
Whether images should be saved in a folder structure based on provided metadata. Defaults to False. |
False
|
transparent_background
|
bool
|
Whether the background inside and outside of figures should be transparent. Defaults to False. |
False
|
**kwargs
|
dict[str, Any]
|
Keyword arguments passed to wrapped function call of |
{}
|
Source code in earthcarekit/plot/figure/swath.py
to_texture
Convert the figure to a texture by removing all axis ticks, labels, annotations, and text.
Source code in earthcarekit/plot/figure/swath.py
add_depol_ratio
add_depol_ratio(
ds_anom,
rolling_w=20,
near_zero_tolerance=2e-07,
smooth=True,
skip_height_above_elevation=300,
cpol_var="mie_attenuated_backscatter",
xpol_var="crosspolar_attenuated_backscatter",
elevation_var=ELEVATION_VAR,
height_var=HEIGHT_VAR,
height_dim=VERTICAL_DIM,
)
Compute depolarization ratio (DPOL = XPOL/CPOL) from attenuated backscatter signals.
This function derives the depol. ratio from cross-polarized (XPOL) and co-polarized (CPOL) attenuated backscatter signals.
Signals below the surface are masked, by default with a vertical margin on 300 meters above elevation to remove potential surface return.
Also, signals are smoothed (or "cleaned") with a rolling mean, and near-zero divisions are suppressed and set to NaN instead.
In the resulting dataset, the ratio curtain and a ratio profile calculated from mean profiles of the full dataset (e.g., mean(XPOL)/mean(CPOL)).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds_anom
|
Dataset
|
ATL_NOM_1B dataset containing cross- and co-polar attenuated backscatter. |
required |
rolling_w
|
int
|
Window size for rolling mean smoothing. Defaults to 20. |
20
|
near_zero_tolerance
|
float
|
Tolerance for masking near-zero |
2e-07
|
smooth
|
bool
|
Whether to apply rolling mean smoothing. Defaults to True. |
True
|
skip_height_above_elevation
|
int
|
Vertical margin above surface elevation to mask in meters. Defaults to 300. |
300
|
cpol_var
|
str
|
Input co-polar variable name. Defaults to "mie_attenuated_backscatter". |
'mie_attenuated_backscatter'
|
xpol_var
|
str
|
Input cross-polar variable name. Defaults to "crosspolar_attenuated_backscatter". |
'crosspolar_attenuated_backscatter'
|
elevation_var
|
str
|
Elevation variable name. Defaults to ELEVATION_VAR. |
ELEVATION_VAR
|
height_var
|
str
|
Height variable name. Defaults to HEIGHT_VAR. |
HEIGHT_VAR
|
height_dim
|
str
|
Height dimension name. Defaults to VERTICAL_DIM. |
VERTICAL_DIM
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: Dataset with added depol. ratio, cleaned signals, and depol. ratio profile from mean profiles. |
Source code in earthcarekit/utils/read/product/level1/atl_nom_1b.py
add_isccp_cloud_type
add_isccp_cloud_type(
ds,
new_var="isccp_cloud_type",
cot_var="cloud_optical_thickness",
cth_var="cloud_top_height",
along_track_dim=ALONG_TRACK_DIM,
across_track_dim=ACROSS_TRACK_DIM,
)
Adds a variable to the dataset containing ISCCP cloud types calculated from cloud optical thickness (COT) and cloud top height (CTH).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
Dataset
|
A MSI_COP_2A dataset. |
required |
new_var
|
str
|
Name of the new ISCCP cloud type variable. Defaults to "isccp_cloud_type". |
'isccp_cloud_type'
|
cot_var
|
str
|
Name of the COT variable in |
'cloud_optical_thickness'
|
cth_var
|
str
|
Name of the CTH variable in |
'cloud_top_height'
|
along_track_dim
|
str
|
Name of the along-track dimension in |
ALONG_TRACK_DIM
|
across_track_dim
|
str
|
Name of the across-track dimension in |
ACROSS_TRACK_DIM
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: The input dataset with added ISCCP cloud type variable. |
References
- International Satellite Cloud Climatology Project (ISCCP). ISCCP Definition of Cloud Types. Retrieved September 25, 2025. https://isccp.giss.nasa.gov/cloudtypes.html
Source code in earthcarekit/utils/read/product/level2a/msi_cop_2a.py
add_potential_temperature
add_potential_temperature(
ds,
temperature_var="temperature_kelvin",
pressure_var="pressure",
new_var="potential_temperature",
)
Computes potential temperature from temperature [K] and pressure [Pa] and adds it as a variable to the dataset (source: https://en.wikipedia.org/wiki/Potential_temperature, accessed: 2026-02-06).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
Dataset
|
Dataset (e.g., AUX_MET_1D) containing temperature [K] and pressure [Pa] data. |
required |
temperature_var
|
str
|
Input temperature variable name. Defaults to "temperature_kelvin". |
'temperature_kelvin'
|
pressure_var
|
str
|
Input pressure variable name. Defaults to "pressure". |
'pressure'
|
new_var
|
str
|
New variable name for potential temperature. Defaults to "potential_temperature". |
'potential_temperature'
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: Dataset with 2 new variables for potential temperature profiles added (kelvin and celsius). |
Source code in earthcarekit/utils/read/product/auxiliary/aux_met_1d.py
add_scattering_ratio
add_scattering_ratio(
ds_anom,
formula,
rolling_w=20,
near_zero_tolerance=2e-07,
smooth=True,
skip_height_above_elevation=300,
cpol_var="mie_attenuated_backscatter",
xpol_var="crosspolar_attenuated_backscatter",
ray_var="rayleigh_attenuated_backscatter",
elevation_var=ELEVATION_VAR,
height_var=HEIGHT_VAR,
height_dim=VERTICAL_DIM,
)
Compute scattering ratio from attenuated backscatter signals given a formula: "x/c", "(c+x)/r", or "(c+x+r)/r".
This function derives the scattering ratio from cross-polarized (XPOL), co-polarized (CPOL) and rayleigh (RAY) attenuated backscatter signals.
Signals below the surface are masked, by default with a vertical margin on 300 meters above elevation to remove potential surface return.
Also, signals are smoothed (or "cleaned") with a rolling mean, and near-zero divisions are suppressed and set to NaN instead.
In the resulting dataset, the ratio curtain and a ratio profile calculated from mean profiles of the full dataset (e.g., mean(XPOL)/mean(CPOL)).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds_anom
|
Dataset
|
ATL_NOM_1B dataset containing the attenuated backscatter signals. |
required |
formula
|
Literal['x/c', '(c+x)/r', '(c+x+r)/r']
|
Formula used to calculate the scattering ratio. |
required |
rolling_w
|
int
|
Window size for rolling mean smoothing. Defaults to 20. |
20
|
near_zero_tolerance
|
float
|
Tolerance for masking near-zero denominators. Defaults to 2e-7. |
2e-07
|
smooth
|
bool
|
Whether to apply rolling mean smoothing. Defaults to True. |
True
|
skip_height_above_elevation
|
int
|
Vertical margin above surface elevation to mask in meters. Defaults to 300. |
300
|
cpol_var
|
str
|
Input co-polar variable name. Defaults to "mie_attenuated_backscatter". |
'mie_attenuated_backscatter'
|
xpol_var
|
str
|
Input cross-polar variable name. Defaults to "crosspolar_attenuated_backscatter". |
'crosspolar_attenuated_backscatter'
|
ray_var
|
str
|
Input rayleigh variable name. Defaults to "rayleigh_attenuated_backscatter". |
'rayleigh_attenuated_backscatter'
|
elevation_var
|
str
|
Elevation variable name. Defaults to ELEVATION_VAR. |
ELEVATION_VAR
|
height_var
|
str
|
Height variable name. Defaults to HEIGHT_VAR. |
HEIGHT_VAR
|
height_dim
|
str
|
Height dimension name. Defaults to VERTICAL_DIM. |
VERTICAL_DIM
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: xr.Dataset: Dataset with added ratio curtain and ratio profile from mean profiles. |
Source code in earthcarekit/utils/read/product/level1/atl_nom_1b.py
39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 | |
compare_bsc_ext_lr_depol
compare_bsc_ext_lr_depol(
input_ec,
input_ground=None,
time_var_ground="time",
height_var_ground="height",
bsc_var_ground=[],
ext_var_ground=[],
lr_var_ground=[],
depol_var_ground=[],
input_ec2=None,
input_ec3=None,
input_ec4=None,
input_ground2=None,
input_ground3=None,
input_ground4=None,
time_var_ground2=None,
height_var_ground2=None,
time_var_ground3=None,
height_var_ground3=None,
time_var_ground4=None,
height_var_ground4=None,
bsc_var_ground2=None,
ext_var_ground2=None,
lr_var_ground2=None,
depol_var_ground2=None,
bsc_var_ground3=None,
ext_var_ground3=None,
lr_var_ground3=None,
depol_var_ground3=None,
bsc_var_ground4=None,
ext_var_ground4=None,
lr_var_ground4=None,
depol_var_ground4=None,
site=None,
radius_km=100.0,
resolution="_low_resolution",
resolution2=None,
resolution3=None,
resolution4=None,
height_range=(0, 30000.0),
selection_height_range=None,
selection_height_range_bsc=None,
selection_height_range_ext=None,
selection_height_range_lr=None,
selection_height_range_depol=None,
value_range_bsc=(0, 8e-06),
value_range_ext=(0, 0.0003),
value_range_lr=(0, 100),
value_range_depol=(0, 0.6),
colors_ec=["ec:red", "ec:orange", "ec:yellow", "ec:purple"],
colors_ground=[
"ec:blue",
"ec:darkblue",
"ec:lightgreen",
"ec:darkgreen",
"ec:purple",
],
linewidth_ec=1.5,
linewidth_ground=1.5,
linestyle_ec="solid",
linestyle_ground="solid",
label_ec=[],
label_ground=[],
alpha=1.0,
show_steps=DEFAULT_PROFILE_SHOW_STEPS,
show_error_ec=False,
show_quality_status=False,
show_rebinned=False,
quality_status_width_scale=1.0,
quality_status_var="quality_status",
quality_status_value_range=None,
to_mega=False,
single_figsize=(5 * CM_AS_INCH, 12 * CM_AS_INCH),
label_bsc="Bsc. coeff.",
label_ext="Ext. coeff.",
label_lr="Lidar ratio",
label_depol="Depol. ratio",
units_bsc="m$^{-1}$ sr$^{-1}$",
units_ext="m$^{-1}$",
units_lr="sr",
units_depol="",
verbose=True,
)
Compares Lidar profiles from up to 3 EarthCARE source dataset an one ground-based dataset by creating plots and statistics dataframe.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
input_ec
|
str | Dataset
|
A opened EarthCARE or file path. |
required |
input_ground
|
str | Dataset
|
A opened ground-based NetCDF dataset or file path (e.g., PollyNET data). |
None
|
time_var_ground
|
str
|
The name of the time variable in the ground-based dataset (e.g., for single profile PollyNET data use |
'time'
|
height_var_ground
|
str
|
The name of the height variable in the ground-based dataset. Defaults to |
'height'
|
bsc_var_ground
|
str | tuple | list[str | tuple]
|
Backscatter variable name in the ground-based dataset.
Multiple variables can be provided as list. Variable errors can be provided as tuples (e.g., |
[]
|
ext_var_ground
|
str | tuple | list[str | tuple]
|
Extinction variable name in the ground-based dataset.
Multiple variables can be provided as list. Variable errors can be provided as tuples (e.g., |
[]
|
lr_var_ground
|
str | tuple | list[str | tuple]
|
Lidar ratio variable name in the ground-based dataset.
Multiple variables can be provided as list. Variable errors can be provided as tuples (e.g., |
[]
|
depol_var_ground
|
str | tuple | list[str | tuple]
|
Depol. ratio variable name in the ground-based dataset.
Multiple variables can be provided as list. Variable errors can be provided as tuples (e.g., |
[]
|
input_ec2
|
str | Dataset
|
An optional seconds EarthCARE dataset to compare. Defaults to None. |
None
|
input_ec3
|
str | Dataset
|
An optional third EarthCARE dataset to compare. Defaults to None. |
None
|
site
|
GroundSite | str | None
|
Ground site or location of the ground-based data as a |
None
|
radius_km
|
float
|
Radius around the ground site. Defaults to 100.0. |
100.0
|
resolution
|
str
|
Sets the used resolution of the EarthCARE data if applicable (e.g., for A-EBD). Defaults to "_low_resolution". |
'_low_resolution'
|
height_range
|
tuple[float, float] | None
|
Height range in meters to restrict the data for plotting. Defaults to (0, 30e3). |
(0, 30000.0)
|
selection_height_range
|
tuple[float, float] | None
|
Height range in meters to select data for statistsics. Defaults to None. |
None
|
selection_height_range_bsc
|
tuple[float, float] | None
|
Height range in meters to select bsc. data for statistsics. Defaults to None (i.e., |
None
|
selection_height_range_ext
|
tuple[float, float] | None
|
Height range in meters to select ext. data for statistsics. Defaults to None (i.e., |
None
|
selection_height_range_lr
|
tuple[float, float] | None
|
Height range in meters to select LR data for statistsics. Defaults to None (i.e., |
None
|
selection_height_range_depol
|
tuple[float, float] | None
|
Height range in meters to select depol. data for statistsics. Defaults to None (i.e., |
None
|
value_range_bsc
|
ValueRangeLike | None
|
Tuple setting minimum and maximum value on x-axis. Defaults to (0, 8e-6). |
(0, 8e-06)
|
value_range_ext
|
ValueRangeLike | None
|
Tuple setting minimum and maximum value on x-axis. Defaults to (0, 3e-4). |
(0, 0.0003)
|
value_range_lr
|
ValueRangeLike | None
|
Tuple setting minimum and maximum value on x-axis. Defaults to (0, 100). |
(0, 100)
|
value_range_depol
|
ValueRangeLike | None
|
Tuple setting minimum and maximum value on x-axis. Defaults to (0, 0.6). |
(0, 0.6)
|
colors_ec
|
list[str]
|
List of colors for the EarthCARE profiles. |
['ec:red', 'ec:orange', 'ec:yellow', 'ec:purple']
|
colors_ground
|
list[str]
|
List of colors for the ground-based profiles. |
['ec:blue', 'ec:darkblue', 'ec:lightgreen', 'ec:darkgreen', 'ec:purple']
|
linewidth_ec
|
Number | list[Number]
|
Value or list of line width for the EarthCARE profiles. Defaults to 1.5. |
1.5
|
linewidth_ground
|
Number | list[Number]
|
Value or list of line width for the ground-based profiles. Defaults to 1.5. |
1.5
|
linestyle_ec
|
Number | list[Number]
|
Value or list of line style for the EarthCARE profiles. Defaults to "solid". |
'solid'
|
linestyle_ground
|
Number | list[Number]
|
Value or list of line style for the ground-based profiles. Defaults to "solid". |
'solid'
|
label_ec
|
list[str]
|
List of legend labels for the EarthCARE profiles. |
[]
|
label_ground
|
list[str]
|
List of legend labels for the ground-based profiles. |
[]
|
alpha
|
float
|
Transparency value for the profile lines (value between 0 and 1). Defaults to 1.0. |
1.0
|
show_steps
|
bool
|
If True, profiles will be plotted as step functions instead of bin centers. |
DEFAULT_PROFILE_SHOW_STEPS
|
show_error_ec
|
bool
|
If True, plot error ribbons for EarthCARE profiles. |
False
|
show_rebinned
|
bool
|
If True, ground-based profiles will be plotted rebinnned to the first EarthCARE profile. Defaults to False. |
False
|
to_mega
|
bool
|
If Ture, converts bsc. and ext. data results (i.e., plot and statistics) to [Mm-1 sr-1] and [Mm-1]. Defaults to False. |
False
|
single_figsize
|
tuple[float, float]
|
2-element tuple setting width and height of the subfigures (i.e., for each profile plot). |
(5 * CM_AS_INCH, 12 * CM_AS_INCH)
|
label_bsc
|
str
|
Label displayed on the backscatter sub-figure. Defaults to "Bsc. coeff.". |
'Bsc. coeff.'
|
label_ext
|
str
|
Label displayed on the extinction sub-figure. Defaults to "Ext. coeff.". |
'Ext. coeff.'
|
label_lr
|
str
|
Label displayed on the lidar ratio sub-figure. Defaults to "Lidar ratio". |
'Lidar ratio'
|
label_depol
|
str
|
Label displayed on the depol sub-figure. Defaults to "Depol. ratio". |
'Depol. ratio'
|
units_bsc
|
str
|
Units displayed on the backscatter sub-figure. Defaults to "m$^{-1}$ sr$^{-1}$". |
'm$^{-1}$ sr$^{-1}$'
|
units_ext
|
str
|
Units displayed on the extinction sub-figure. Defaults to "m$^{-1}$". |
'm$^{-1}$'
|
units_lr
|
str
|
Units displayed on the lidar ratio sub-figure. Defaults to "sr". |
'sr'
|
units_depol
|
str
|
Units displayed on the depol sub-figure. Defaults to "". |
''
|
verbose
|
bool
|
Whether logs about processing steps appear in the console. Defaults to True. |
True
|
Returns:
| Name | Type | Description |
|---|---|---|
results |
_CompareBscExtLRDepolResults
|
An object containing the plot and statistical results.
- |
Source code in earthcarekit/calval/_compare_bsc_ext_lr_depol.py
557 558 559 560 561 562 563 564 565 566 567 568 569 570 571 572 573 574 575 576 577 578 579 580 581 582 583 584 585 586 587 588 589 590 591 592 593 594 595 596 597 598 599 600 601 602 603 604 605 606 607 608 609 610 611 612 613 614 615 616 617 618 619 620 621 622 623 624 625 626 627 628 629 630 631 632 633 634 635 636 637 638 639 640 641 642 643 644 645 646 647 648 649 650 651 652 653 654 655 656 657 658 659 660 661 662 663 664 665 666 667 668 669 670 671 672 673 674 675 676 677 678 679 680 681 682 683 684 685 686 687 688 689 690 691 692 693 694 695 696 697 698 699 700 701 702 703 704 705 706 707 708 709 710 711 712 713 714 715 716 717 718 719 720 721 722 723 724 725 726 727 728 729 730 731 732 733 734 735 736 737 738 739 740 741 742 743 744 745 746 747 748 749 750 751 752 753 754 755 756 757 758 759 760 761 762 763 764 765 766 767 768 769 770 771 772 773 774 775 776 777 778 779 780 781 782 783 784 785 786 787 788 789 790 791 792 793 794 795 796 797 798 799 800 801 802 803 804 805 806 807 808 809 810 811 812 813 814 815 816 817 818 819 820 821 822 823 824 825 826 827 828 829 830 831 832 833 834 835 836 837 838 839 840 841 842 843 844 845 846 847 848 849 850 851 852 853 854 855 856 857 858 859 860 861 862 863 864 865 866 867 868 869 870 871 872 873 874 875 876 877 878 879 880 881 882 883 884 885 886 887 888 889 890 891 892 893 894 895 896 897 898 899 900 901 902 903 904 905 906 907 908 909 910 911 912 913 914 915 916 917 918 919 920 921 922 923 924 925 926 927 928 929 930 931 932 933 934 935 936 937 938 939 940 941 942 943 944 945 946 947 948 949 950 951 952 953 954 955 956 957 958 959 960 961 962 963 964 965 966 967 968 969 970 971 972 973 974 975 976 977 978 979 980 981 982 983 984 985 986 987 988 989 990 991 992 993 994 995 996 997 998 999 1000 1001 1002 1003 1004 1005 1006 1007 1008 1009 1010 1011 1012 1013 1014 1015 1016 1017 1018 1019 1020 1021 1022 1023 1024 1025 1026 1027 1028 1029 1030 1031 1032 1033 | |
create_column_figure_layout
create_column_figure_layout(
ncols, single_figsize=(3, 8), margin=0.0, height_scale=1.0, width_scale=1.0
)
Creates a figure with multiple subfigures arranged as columns in a single row, each containing one Axes.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ncols
|
int
|
Number of subfigures (columns) to create. |
required |
single_figsize
|
tuple[float, float]
|
Size (width, height) of each individual subfigure. Defaults to (3, 8). |
(3, 8)
|
Returns:
| Type | Description |
|---|---|
FigureLayoutColumns
|
tuple[Figure, list[Axes]]: The parent figure and a list of Axes objects, one for each subfigure. |
Source code in earthcarekit/plot/figure/multi_panel/simple_columns.py
create_multi_figure_layout
create_multi_figure_layout(
rows,
zoom_rows=None,
profile_rows=None,
map_rows=None,
wspace=1.2,
hspace=1.2,
wmain=FIGURE_WIDTH_CURTAIN,
hrow=FIGURE_HEIGHT_CURTAIN,
hswath=FIGURE_HEIGHT_SWATH,
hline=FIGURE_HEIGHT_LINE,
wprofile=FIGURE_WIDTH_PROFILE,
wmap=FIGURE_MAP_WIDTH,
wzoom=FIGURE_WIDTH_CURTAIN / 3.0,
)
Creates a complex figure layout with columns for map, main, zoom, and profile panels (in that order from left to right).
Each panel column can have a custom sequence of figure types (e.g., row heights), and the layout supports both uniform and per-gap horizontal/vertical spacing.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
main_rows
|
Sequence[FigureType | int]
|
List of figure types for the rows of the main column. |
required |
zoom_rows
|
Sequence[FigureType | int]
|
List of figure types for the rows in the optional zoom column. |
None
|
profile_rows
|
Sequence[FigureType | int]
|
List of figure types for the rows in the optional profile column. |
None
|
map_rows
|
Sequence[FigureType | int]
|
List of figure types for the rows in the optional map column. |
None
|
wspace
|
float | Sequence[float]
|
Horizontal spacing between columns. Can be a single value or list defining spacing before, between, and after columns. |
1.2
|
hspace
|
float | Sequence[float]
|
Vertical spacing between rows. Similar behavior as |
1.2
|
wmain
|
float
|
Width of the main column. Default is |
FIGURE_WIDTH_CURTAIN
|
hrow
|
float
|
Height of a standard row. Default is |
FIGURE_HEIGHT_CURTAIN
|
hswath
|
float
|
Height of a |
FIGURE_HEIGHT_SWATH
|
wprofile
|
float
|
Width of the profile column. |
FIGURE_WIDTH_PROFILE
|
wmap
|
float
|
Width of the map column. |
FIGURE_MAP_WIDTH
|
wzoom
|
float
|
Width of the zoom column. |
FIGURE_WIDTH_CURTAIN / 3.0
|
Returns:
| Name | Type | Description |
|---|---|---|
tuple |
FigureLayoutMapMainZoomProfile
|
A tuple containing: - Figure: The matplotlib figure object. - Sequence[Axes]: Axes for map panels (may be empty). - Sequence[Axes]: Axes for main panels. - Sequence[Axes]: Axes for zoom panels (may be empty). - Sequence[Axes]: Axes for profile panels (may be empty). |
Raises:
| Type | Description |
|---|---|
ValueError
|
If the provided spacing sequences are of invalid length. |
TypeError
|
If spacing arguments are of unsupported types. |
Source code in earthcarekit/plot/figure/multi_panel/map_main_zoom_profile_figure.py
31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 | |
ecdownload
ecdownload(
file_type,
baseline=None,
orbit_number=None,
start_orbit_number=None,
end_orbit_number=None,
frame_id=None,
orbit_and_frame=None,
start_orbit_and_frame=None,
end_orbit_and_frame=None,
timestamps=None,
start_time=None,
end_time=None,
radius_search=None,
bounding_box=None,
path_to_config=None,
path_to_data=None,
is_log=False,
is_debug=False,
is_download=True,
is_overwrite=False,
is_unzip=True,
is_delete=True,
is_create_subdirs=True,
is_export_results=False,
idx_selected_input=None,
is_organize_data=False,
is_include_header=None,
is_reversed_order=False,
return_results=False,
verbose=True,
check_product_availability=False,
)
EarthCARE Download Tool: Search for and download EarthCARE products from a ESA data distribution platform (OADS or MAAP).
The execution of this tool is divided into two parts:
- First, based on provided arguments search request will be send via the OpenSearch API of the ESA MAAP catalogue.
- Second, the resulting list of products is then downloaded from the configures download backend (OADS or MAAP). See:
- MAAP: portal.maap.eo.esa.int/earthcare
- OADS L1: ec-pdgs-dissemination1.eo.esa.int
- OADS L2: ec-pdgs-dissemination2.eo.esa.int
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
file_type
|
str | list[str]
|
Name(s) of EarthCARE product(s) to search for (e.g., "ATL_NOM_1B", "ANOM", or "A-NOM"). Note: Input string evaluation is not case sensitive. Also, product version may also be selected by adding a colon and the two-letter processor baseline after the name (e.g., "ANOM:BA"). |
required |
baseline
|
str | None
|
Two-letter processor baseline used as default for all given |
None
|
orbit_number
|
int | list[int] | None
|
Specific orbit number(s) to search for (e.g., 981 or [1000, 5000, ...]). Defaults to None. |
None
|
start_orbit_number
|
int | None
|
The lower limit of orbit numbers to search for (e.g., 5000). Defaults to None. |
None
|
end_orbit_number
|
int | None
|
The upper limit of orbit numbers to search for (e.g., 5003). Defaults to None. |
None
|
frame_id
|
str | list[str] | None
|
Frame ID letter(s) to search for (i.e., letters A to H). Defaults to None. |
None
|
orbit_and_frame
|
str | list[str] | None
|
Orbit and frame string(s) to search for (e.g., "01234F" or ["1000A", "5000C", ...]). Defaults to None. |
None
|
start_orbit_and_frame
|
str | None
|
The lower limit of orbit and frames to search for (e.g., "05000D"). Defaults to None. |
None
|
end_orbit_and_frame
|
str | None
|
The upper limit of orbit and frames to search for (e.g., "05003C"). Defaults to None. |
None
|
timestamps
|
str | list[str] | None
|
Search for data containing specific timestamp(s) (e.g. "2024-07-31 13:45" or "20240731T134500Z"). Defaults to None. |
None
|
start_time
|
str | None
|
The lower time limit for the search. Defaults to None. |
None
|
end_time
|
str | None
|
The upper time limit for the search. Defaults to None. |
None
|
radius_search
|
tuple[RadiusMetersFloat, LatFloat, LonFloat] | list | None
|
A tuple containing a radius (meters) and a lat/lon point to perform a geo radius search (e.g., 25000 51.35 12.43, i.e.,
|
None
|
bounding_box
|
tuple[LatSFloat, LonWFloat, LatNFloat, LonEFloat] | list | None
|
A tuple containing the extent for a bounding box geo search (e.g., [14.9, 37.7, 14.99, 37.78],
i.e., |
None
|
path_to_config
|
str | None
|
If provided, uses given config file instead of the default config. Defaults to None. |
None
|
path_to_data
|
str | None
|
If provided, downloads data to the given folder instead of the one defined in the config file. Defaults to None. |
None
|
is_log
|
bool
|
If True, creates a log file in a |
False
|
is_debug
|
bool
|
If True, shows debug logs in the console. Defaults to False. |
False
|
is_download
|
bool
|
If False, skips download part, but still performs search requests via the data dissemination platform API. Defaults to True. |
True
|
is_overwrite
|
bool
|
If True, downloads and overwrites files that already exist in the data directory instead of skipping them. Defaults to False. |
False
|
is_unzip
|
bool
|
If False, skips file extraction for downloaded archives. Defaults to True. |
True
|
is_delete
|
bool
|
If True, deletes downloaded archives after extraction (i.e., does not delete non-extracted archives). Defaults to True. |
True
|
is_create_subdirs
|
bool
|
If True, places downloaded files in a sub-directory structure according to the template defined in the config file. Defaults to True. |
True
|
is_export_results
|
bool
|
If True, creates a text file in the current working directory listing all search results. Defaults to False. |
False
|
idx_selected_input
|
int | None
|
A number matching an index in the list of found files. If provided, only this single file will be downloaded. Defaults to None. |
None
|
is_organize_data
|
bool
|
If True, does not search or download any data. Defaults to False. |
False
|
is_include_header
|
bool | None
|
If True, the full archive is downloaded containing both HDF5 data file ( Caution This option only applies to MAAP. OADS will always download the full archive. |
None
|
is_reversed_order
|
bool
|
If True, downloads data products in reversed order (from the latest to the earliest). Defaults to False. |
False
|
return_results
|
bool
|
If True, returns the search results as a |
False
|
verbose
|
bool
|
If False, does not print logs to the console and does not create log file. Defaults to True. |
True
|
check_product_availability
|
bool
|
If True, sends extra request to the download backend checking the list of available products per data collection. If False, uses internally stored lists of available products, significantly reducing execution time (but might fail in case of backend changes). Defaults to False. |
False
|
Returns:
| Name | Type | Description |
|---|---|---|
results |
ProductDataFrame | None
|
If |
Source code in earthcarekit/download/main.py
40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 | |
ecquicklook
ecquicklook(
ds,
vars=None,
show_maps=True,
show_zoom=False,
show_profile=True,
site=None,
radius_km=100.0,
time_range=None,
height_range=None,
ds_tropopause=None,
ds_elevation=None,
ds_temperature=None,
resolution="medium",
ds2=None,
ds_xmet=None,
logger=None,
log_msg_prefix="",
selection_max_time_margin=None,
show_steps=DEFAULT_PROFILE_SHOW_STEPS,
mode="fast",
map_style="blue_marble",
curtain_kwargs={},
map_kwargs={},
profile_kwargs={},
)
Generate a preview visualization of an EarthCARE dataset with optional maps, zoomed views, and profiles.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
Dataset | str
|
EarthCARE dataset or path. |
required |
vars
|
(str | list[str] | None, otional)
|
List of variable to plot. Automatically sets product-specific default list of variables if None. |
None
|
show_maps
|
bool
|
Whether to include map view. Dafaults to True. |
True
|
show_zoom
|
bool
|
Whether to show an additional column of zoomed plots. Defaults to False. |
False
|
show_profile
|
bool
|
Whether to include vertical profile plots. Dfaults to True. |
True
|
site
|
GroundSite | str | None
|
Ground site object or name identifier. |
None
|
radius_km
|
float
|
Search radius around site in kilometers. Defaults to 100. |
100.0
|
time_range
|
TimeRangeLike | None
|
Time range filter. |
None
|
height_range
|
DistanceRangeNoneLike | None
|
Height range in meters. Defaults to None. |
None
|
ds_tropopause
|
Dataset | str | None
|
Optional dataset or path containing tropopause data to add it to the plot. |
None
|
ds_elevation
|
Dataset | str | None
|
Optional dataset or path containing elevation data to add it to the plot. |
None
|
ds_temperature
|
Dataset | str | None
|
Optional dataset or path containing temperature data to add it to the plot. |
None
|
resolution
|
Literal['low', 'medium', 'high', 'l', 'm', 'h']
|
Resolution of A-PRO data. Defaults to "low". |
'medium'
|
ds2
|
Dataset | str | None
|
Secondary dataset required for certain product quicklook (e.g., A-LAY products need A-NOM or A-EBD to serve as background curtain plots). |
None
|
ds_xmet
|
Dataset | str | None
|
Optional auxiliary meteorological dataset used to plot tropopause, elevation and temperature from. |
None
|
logger
|
Logger
|
Logger instance for output messages. |
None
|
log_msg_prefix
|
str
|
Prefix for log messages. |
''
|
selection_max_time_margin
|
TimedeltaLike | Sequence[TimedeltaLike] | None
|
Allowed time difference for selection. |
None
|
show_steps
|
bool
|
Whether to plot profiles as height bin step functions or instead plot only the line through bin centers. Defaults to True. |
DEFAULT_PROFILE_SHOW_STEPS
|
mode
|
Literal['fast', 'exact']
|
Processing mode. |
'fast'
|
map_style
|
str | Literal['none', 'stock_img', 'gray', 'osm', 'satellite', 'mtg', 'msg', 'blue_marble', 'land_ocean', 'land_ocean_lakes_rivers'] | None
|
Style of the background in the secondary/zoomed map. Defaults to "blue_marble". |
'blue_marble'
|
Returns:
| Name | Type | Description |
|---|---|---|
_QuicklookResults |
QuicklookFigure
|
Object containing figures and metadata. |
Source code in earthcarekit/plot/quicklook/_quicklook.py
90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 244 245 246 247 248 249 250 251 252 253 254 255 256 257 258 259 | |
ecquicklook_deep_convection
ecquicklook_deep_convection(
mrgr,
cfmr,
ccd,
aebd,
xmet=None,
height_range=(-250, 20000.0),
time_range=None,
info_text_loc=None,
trim_to_frame=False,
)
Creates a 4 panel quicklook of a storm or deep convective event, displaying:
- 1st row: RGB image from MSI_RGR_1C
- 2nd row: Radar reflectivity from CPR_FMR_2A
- 3rd row: Doppler velocity from CPR_CD__2A
- 4th row: Total attenuated backscatter from ATL_EBD_2A
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds_mrgr
|
Dataset
|
The MSI_RGR_1C product filepath or dataset. |
required |
ds_cfmr
|
Dataset
|
The CPR_FMR_2A product filepath or dataset. |
required |
ds_ccd
|
Dataset
|
The CPR_CD__2A product filepath or dataset. |
required |
ds_aebd
|
Dataset
|
The ATL_EBD_2A product filepath or dataset. |
required |
ds_xmet
|
Dataset | None
|
The AUX_MET_1D product filepath or dataset. If given, temperature contour lines will be added to the plots. Defaults to None. |
required |
height_range
|
DistanceRangeLike | None
|
A height range (i.e., min, max) in meters. Defaults to (-250, 20e3). |
(-250, 20000.0)
|
time_range
|
TimeRangeLike | None
|
A time range to filter the displayed data. Defaults to None. |
None
|
info_text_loc
|
str | None
|
The positioning of the orbt, frame and product info text (e.g., "upper right"). Defaults to None. |
None
|
trim_to_frame
|
bool
|
Wether the read products should be trimmed to the EarthCARE frame bounds. |
False
|
Returns:
| Name | Type | Description |
|---|---|---|
QuicklookFigure |
QuicklookFigure
|
The quicklook object. |
Examples:
import earthcarekit as eck
df = eck.search_product(
file_type=["mrgr", "cfmr", "ccd", "aebd", "xmet"],
orbit_and_frame="07590D",
).filter_latest()
fp_mrgr = df.filter_file_type("mrgr").filepath[-1]
fp_cfmr = df.filter_file_type("cfmr").filepath[-1]
fp_ccd = df.filter_file_type("ccd").filepath[-1]
fp_aebd = df.filter_file_type("aebd").filepath[-1]
fp_xmet = df.filter_file_type("xmet").filepath[-1]
ql = eck.ecquicklook_deep_convection(
mrgr=fp_mrgr,
cfmr=fp_cfmr,
ccd=fp_ccd,
aebd=fp_aebd,
xmet=fp_xmet,
time_range=("2025-09-28T18:27:10", None),
info_text_loc="upper left",
)
Source code in earthcarekit/plot/quicklook/_quicklook_deep_convection.py
18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 | |
ecquicklook_psc
ecquicklook_psc(
anom,
xmet=None,
zoom_at=0.5,
height_range=(0, 40000.0),
time_range=None,
info_text_loc=None,
)
Creates a two-column multi-panel quicklook of a PSC event, displaying:
- 1st column: Two maps showing the EarthCARE track.
- 2nd column: Three rows showing co- and cross-polar attenuated backscatter and the calculated depolarization ratio.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
anom
|
str | Sequence[str] | Dataset
|
The ATL_NOM_1B product filepath(s) or dataset(s). |
required |
xmet
|
str | Sequence[str] | Dataset | None
|
The AUX_MET_1D product filepath(s) or dataset(s). If given, temperature contour lines will be added to the plots. Defaults to None. |
None
|
zoom_at
|
float | None
|
In case two frames are given, selects only a zoomed-in portion of the frames around this fractional index (0 -> only 1st frame, 0.5 -> half of end of 1st and half of beginning of 2nd frame, 1 -> only 2nd frame). Defaults to 0.5. |
0.5
|
height_range
|
DistanceRangeLike | None
|
description. Defaults to (0, 40e3). |
(0, 40000.0)
|
time_range
|
TimeRangeLike | None
|
A time range to filter the displayed data. Defaults to None. |
None
|
info_text_loc
|
str | None
|
The positioning of the orbt, frame and product info text (e.g., "upper right"). Defaults to None. |
None
|
Raises:
| Type | Description |
|---|---|
ValueError
|
If none or more than 2 frames are given. |
ValueError
|
If given number X-MET files does not match number of A-NOM files. |
Returns:
| Name | Type | Description |
|---|---|---|
QuicklookFigure |
QuicklookFigure
|
The quicklook object. |
Examples:
import earthcarekit as eck
df = eck.search_product(
file_type=["anom", "xmet"],
orbit_and_frame=["3579B", "3579C"],
).filter_latest()
fps_anom = df.filter_file_type("anom").filepath
fps_xmet = df.filter_file_type("xmet").filepath
ql = eck.ecquicklook_psc(
anom=fps_anom,
xmet=fps_xmet,
)
Source code in earthcarekit/plot/quicklook/_quicklook_psc.py
21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 225 226 227 228 229 230 231 232 233 234 235 236 237 238 239 240 241 242 243 | |
geodesic
Calculates the geodesic distances between points on Earth (i.e. WSG 84 ellipsoid) using Vincenty's inverse method.
Supports single or sequences of coordiates.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
a
|
ArrayLike
|
Coordinates [lat, lon] or array of shape (N, 2), in decimal degrees. |
required |
b
|
ArrayLike
|
Second coordinates, same format/shape as |
required |
units
|
str
|
Output units, "km" (default) or "m". |
'km'
|
tolerance
|
float
|
Convergence threshold in radians. Default is 1e-12. |
1e-12
|
max_iterations
|
int
|
Maximum iterations before failure. Default is 10. |
10
|
Returns:
| Type | Description |
|---|---|
float64 | NDArray[float64]
|
float or np.ndarray: The geodesic distance or distances between the point in |
Raises:
| Type | Description |
|---|---|
ValueError
|
If input shapes are incompatible or units are invalid. |
Note
Uses WGS84 (a=6378137.0 m, f=1/298.257223563). May fail for nearly antipodal points.
Examples:
>>> geodesic([51.352757, 12.43392], [38.559, 68.856])
4548.675334434374
>>> geodesic([0,0], [[0,0], [10,0], [20,0]])
array([ 0. , 1105.85483324, 2212.36625417])
>>> geodesic([[0,0], [10,0], [20,0]], [[0,0], [10,0], [20,0]])
array([0., 0., 0.])
References
- Vincenty, T. (1975). Direct and Inverse Solutions of Geodesics on the Ellipsoid with application of nested equations. Survey Review, 23(176), 88-93. https://doi.org/10.1179/sre.1975.23.176.88
Source code in earthcarekit/utils/geo/distance/_vincenty.py
7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 | |
get_cmap
Return a color map given by cmap.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
cmap
|
str | Colormap | list | None
|
|
required |
Returns:
cmap (Cmap):
A color map matching the given cmap.
Source code in earthcarekit/plot/color/colormap/colormap.py
get_config
Returns the default or a given earthcarekit config object.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
c
|
str | ECKConfig | None
|
A path to a config file (.toml) or None. If None, returns the default config. Defaults to None. |
None
|
Returns:
| Name | Type | Description |
|---|---|---|
ECKConfig |
ECKConfig
|
A config object. |
Source code in earthcarekit/utils/config.py
get_coord_between
Interpolates between two coordinates by fraction f (0 to 1).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
coord1
|
ArrayLike
|
The first lat/lon point. |
required |
coord2
|
ArrayLike
|
The second lat/lon point. |
required |
f
|
float
|
A fractional value between 0 and 1. Defaults to 0.5, i.e., the mid point between coord1 and coord2. |
0.5
|
Returns:
| Name | Type | Description |
|---|---|---|
NDArray |
NDArray
|
A 2-element |
Source code in earthcarekit/utils/geo/interpolate.py
get_coords
Takes a xarray.Dataset and returns the lat/lon coordinates as a numpy array.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
lat_var
|
str
|
Name of the latitude variable. Defaults to TRACK_LAT_VAR. |
TRACK_LAT_VAR
|
lon_var
|
str
|
Name of the longitude variable. Defaults to TRACK_LON_VAR. |
TRACK_LON_VAR
|
flatten
|
bool
|
If True, the coordinates will be flattened to a 2D array
|
False
|
Returns:
| Type | Description |
|---|---|
NDArray
|
numpy.array: The extracted lat/lon coordinates. |
Source code in earthcarekit/utils/geo/coordinates.py
get_ground_site
Retruns ground site data based on name and raises ValueError if no matching ground site is found and TypeError.
Source code in earthcarekit/utils/ground_sites.py
get_overpass_info
get_overpass_info(
ds,
site,
radius_km=100.0,
*,
time_var=TIME_VAR,
lat_var=TRACK_LAT_VAR,
lon_var=TRACK_LON_VAR,
along_track_dim=ALONG_TRACK_DIM
)
Extract details about an overpass, including duration, distance, time, closest index, etc.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
str | Dataset
|
Path to or instance of a dataset containing along-track satellite data. |
required |
site
|
GroundSite | str
|
Site name or object over which the satellite is passing. |
required |
radius_km
|
float | int
|
Radius to look for an overpass in kilometers. Defaults to 100. |
100.0
|
time_var
|
str
|
Name of the dataset variable containing time data. Defaults to "time". |
TIME_VAR
|
lat_var
|
str
|
Name of the dataset variable containing latitude data. Defaults to "latitude". |
TRACK_LAT_VAR
|
lon_var
|
str
|
Name of the dataset variable containing longitude data. Defaults to "longitude". |
TRACK_LON_VAR
|
along_track_dim
|
str
|
Name of the along-track or temporal dataset dimension. Defaults to "along_track". |
ALONG_TRACK_DIM
|
Raises:
| Type | Description |
|---|---|
TypeError
|
If |
Returns:
| Name | Type | Description |
|---|---|---|
OverpassInfo |
OverpassInfo
|
description |
Source code in earthcarekit/utils/overpass.py
get_product_info
Gather all info contained in the EarthCARE product's file path.
Source code in earthcarekit/utils/read/product/file_info/product_info.py
89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 | |
get_product_infos
Extracts product metadata from EarthCARE product file paths (e.g. file_type, orbit_number, frame_id, baseline, ...).
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
filepaths
|
str | list[str] | NDArray | DataFrame | Dataset
|
Input sources for EarthCARE product files. Can be one of
- |
required |
**kwargs
|
Additional arguments passed to |
required |
Returns:
| Name | Type | Description |
|---|---|---|
ProductDataFrame |
ProductDataFrame
|
A dataframe containing extracted product information. |
Source code in earthcarekit/utils/read/product/file_info/product_info.py
haversine
Calculates the great-circle (spherical) distance between pairs of latitude/longitude coordinates using the haversine formula.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
a
|
ArrayLike
|
An array-like object of shape (..., 2) containing latitude and longitude coordinates in degrees. The last dimension must be 2: (lat, lon). |
required |
b
|
ArrayLike
|
An array-like object of the same shape as |
required |
units
|
Literal['m', 'km']
|
Unit of the output distance. Must be either "km" for kilometers or "m" for meters. Defaults to "km". |
'km'
|
radius
|
float
|
Radius of the sphere to use for distance calculation.
Defaults to MEAN_EARTH_RADIUS_METERS (based on WSG 84 ellipsoid: ~6371008.77 meters).
Note: If |
required |
Returns:
| Type | Description |
|---|---|
|
np.ndarray: Array of great-circle distances between |
Raises:
| Type | Description |
|---|---|
ValueError
|
If the shapes of |
Examples:
>>> haversine([51.352757, 12.43392], [38.559, 68.856])
4537.564747442274
>>> haversine([0,0], [[0,0], [10,0], [20,0]])
array([ 0. , 1111.95079735, 2223.90159469])
>>> haversine([[0,0], [10,0], [20,0]], [[0,0], [10,0], [20,0]])
array([0., 0., 0.])
Source code in earthcarekit/utils/geo/distance/_haversine.py
perform_anom_depol_statistics
perform_anom_depol_statistics(
ds_anom,
selection_height_range,
is_rayleigh_corrected=False,
rayleigh_correction_factor=0.004,
**kwargs
)
Calculate depolarization statistics and uncertainties within a height range.
This function adds the depol. ratio (DPOL) calculated from co- (CPOL) and cross-polarized (XPOL)
attenuated backscatter to the dataset (ATL_NOM_1B) and computes related statistics.
Mean values and standard deviations are calculated for CPOL, XPOL, and DPOL within the selected
height range. Variability is separated into vertical and temporal components. Errors of DPOL are derived
using error propagation for the XPOL/CPOL ratio.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds_anom
|
Dataset
|
ATL_NOM_1B dataset with cross- and co-polar signals. |
required |
selection_height_range
|
DistanceRangeLike
|
Height range for statistics. |
required |
is_rayleight_corrected
|
bool
|
If True, the mean cross-polar profile is corrected by substracting the mean rayleigh profile scaled by a correction factor. Defaults to False. |
required |
rayleigh_correction_factor
|
float
|
The scaling factor used when |
0.004
|
Returns:
| Name | Type | Description |
|---|---|---|
_ANOMDepolCalculationResults |
_ANOMDepolCalculationResults
|
Results container with
|
Example
import earthcarekit as eck
ft = "ANOM"
oaf = "01508B"
site = "dushanbe"
radius_km = 100
sel_hrange = (1e3, 4e3)
# # Optionally, download required data
# eck.ecdownload(file_type=ft, orbit_and_frame=oaf)
df = eck.search_product(file_type=ft, orbit_and_frame=oaf)
fp = df.filepath[-1]
with eck.read_any(fp) as ds:
ds = eck.filter_radius(ds, radius_km=radius_km, site=site)
results = eck.perform_anom_depol_statistics(ds, sel_hrange)
results.print() # prints statistics
# # Optionally, save statistics as CSV file
# results.stats.to_csv("./stats.csv")
# # Optionally, save profile figure as PNG file
# fig = results.plot(height_range=(0, 10e3))
# eck.save_plot(fig, filepath="./depol_profile.png")
Source code in earthcarekit/calval/_perform_anom_depol_statistics.py
94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 | |
plot_line_between_figures
plot_line_between_figures(
ax1,
ax2,
point1,
point2=None,
color="ec:red",
linestyle="dashed",
linewidth=2,
alpha=0.3,
capstyle="butt",
zorder=-20,
**kwargs
)
Draws a line connecting a point in one subfigure (ax1) to a point in another (ax2).
Source code in earthcarekit/plot/figure/_plot_line_between_figures.py
read_any
Reads various input types and returns an xarray.Dataset.
This function can read
- EarthCARE product files (
.h5) - NetCDF files (
.nc) - Manually processed PollyXT output files (
.txt)
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
input
|
str | Dataset
|
File path or existing Dataset. |
required |
**kwargs
|
Additional keyword arguments for specific readers. |
{}
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: Opened dataset. |
Raises:
| Type | Description |
|---|---|
ValueError
|
If the file type is not supported. |
TypeError
|
If the input type is invalid. |
Source code in earthcarekit/utils/read/_read_any.py
read_header_data
Opens the product header groups of a EarthCARE file as a xarray.Dataset.
Source code in earthcarekit/utils/read/product/header_group.py
read_nc
Returns an xarray.Dataset from a Dataset or NetCDF file path, optionally loaded into memory.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
input
|
Dataset or str
|
Path to a NetCDF file. If a already opened |
required |
modify
|
bool
|
If True, default modifications to the opened dataset will be applied (e.g., converting heights in Polly data from height a.g.l. to height above mean sea level). |
True
|
in_memory
|
bool
|
If True, ensures the dataset is fully loaded into memory. Defaults to False. |
False
|
**kwargs
|
Key-word arguments passed to |
{}
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xarray.Dataset: The resulting dataset. |
Raises:
| Type | Description |
|---|---|
TypeError
|
If input is not a Dataset or string. |
Source code in earthcarekit/utils/read/_read_nc.py
read_polly
Reads manually processed PollyXT output text files as xarray.Dataset or returns an already open one.
Source code in earthcarekit/utils/read/_read_polly.py
read_product
read_product(
input,
trim_to_frame=True,
modify=DEFAULT_READ_EC_PRODUCT_MODIFY,
header=DEFAULT_READ_EC_PRODUCT_HEADER,
meta=DEFAULT_READ_EC_PRODUCT_META,
ensure_nans=DEFAULT_READ_EC_PRODUCT_ENSURE_NANS,
in_memory=False,
**kwargs
)
Returns an xarray.Dataset from a Dataset or EarthCARE file path, optionally loaded into memory.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
input
|
str or Dataset
|
Path to a EarthCARE file. If a |
required |
trim_to_frame
|
bool
|
Whether to trim the dataset to latitude frame bounds. Defaults to True. |
True
|
modify
|
bool
|
If True, default modifications to the opened dataset will be applied (e.g., renaming dimension corresponding to height to "vertical"). Defaults to True. |
DEFAULT_READ_EC_PRODUCT_MODIFY
|
header
|
bool
|
If True, all header data will be included in the dataframe. Defaults to False. |
DEFAULT_READ_EC_PRODUCT_HEADER
|
meta
|
bool
|
If True, select meta data from header (like orbit number and frame ID) will be included in the dataframe. Defaults to True. |
DEFAULT_READ_EC_PRODUCT_META
|
ensure_nans
|
bool
|
If True, ensures that _FillValues are set to NaNs even if encoding of _FillValues or dtype is missing. Be aware, if True increases reading time. Defaults to False. |
DEFAULT_READ_EC_PRODUCT_ENSURE_NANS
|
in_memory
|
bool
|
If True, ensures the dataset is fully loaded into memory. Defaults to False. |
False
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xarray.Dataset: The resulting dataset. |
Raises:
| Type | Description |
|---|---|
TypeError
|
If input is not a Dataset or string. |
Source code in earthcarekit/utils/read/product/_generic.py
read_products
read_products(
filepaths,
zoom_at=None,
along_track_dim=ALONG_TRACK_DIM,
func=None,
func_inputs=None,
max_num_files=8,
coarsen=True,
)
Read and concatenate a sequence of EarthCARE frames into a single xarray Dataset.
By default, the dataset is coarsened according to the number of input frames (e.g.,
combining 3 products averages every 3 profiles, so the along-track dimension remains
comparable to a single product). Optionally applies a processing function to each
frame and zooms in on a specific region (defined by zoom_at) without coarsening.
Coarsening can also be turned of but might case memory issues.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
filepaths
|
Sequence[str] or DataFrame
|
EarthCARE product file paths as a list or a DataFrame with metadata
including |
required |
zoom_at
|
float
|
If set, selects only a zoomed-in portion of the frames around this fractional index. Defaults to None. |
None
|
along_track_dim
|
str
|
Name of the dimension to concatenate along. Defaults to ALONG_TRACK_DIM. |
ALONG_TRACK_DIM
|
func
|
Callable
|
Function to apply to each frame after loading. Defaults to None. |
None
|
func_inputs
|
Sequence[dict]
|
Optional per-frame arguments to pass to |
None
|
max_num_files
|
int
|
Max. number of files that are allowed to be loaded at once.
A |
8
|
coarsen
|
bool
|
If Ture, read data sets are coarened depending on the number given of files. Only aplicable when not zooming. Defaults to Ture. |
True
|
Returns:
| Name | Type | Description |
|---|---|---|
Dataset |
Dataset
|
Concatenated dataset with all frames along |
Source code in earthcarekit/utils/read/product/_concat.py
25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 | |
read_science_data
Opens the science data of a EarthCARE file as a xarray.Dataset.
Source code in earthcarekit/utils/read/product/science_group.py
rebin_msi_to_jsg
rebin_msi_to_jsg(
ds_msi,
ds_xjsg,
vars=None,
k=4,
eps=1e-12,
lat_var=SWATH_LAT_VAR,
lon_var=SWATH_LON_VAR,
time_var=TIME_VAR,
along_track_dim=ALONG_TRACK_DIM,
across_track_dim=ACROSS_TRACK_DIM,
lat_var_xjsg=SWATH_LAT_VAR,
lon_var_xjsg=SWATH_LON_VAR,
time_var_xjsg=TIME_VAR,
along_track_dim_xjsg=ALONG_TRACK_DIM,
across_track_dim_xjsg=ACROSS_TRACK_DIM,
)
Rebins variables from an MSI product dataset onto the geo-spacial lat/lon grid given by the related AUX_JSG_1D dataset.
This function interpolates selected variables from ds_msi onto the JSG grid from ds_xjsg
using quick kd-tree nearest-neighbor search with scipy.spatial.cKDTree followed
by averaging the k-nearest points using inverse distance weighting. The resulting dataframe
matches the along- and across-track resolution of ds_xjsg.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds_msi
|
Dataset | str
|
The source MSI dataset (e.g., MSI_RGR_1C, MSI_COP_2A, ...). |
required |
ds_xjsg
|
Dataset | str
|
The target XJSG dataset. |
required |
vars
|
list[str] | None
|
List of variable names from |
None
|
k
|
int
|
Number of nearest geo-spacial neighbors to include in the kd-tree search. Defaults to 4. |
4
|
eps
|
float
|
Numerical threshold to avoid division by zero in distance calculations during the kd-tree search. Defaults to 1e-12. |
1e-12
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: The MSI dataset with variables rebinned to the JSG grid. |
Source code in earthcarekit/utils/read/product/_rebin_msi_to_jsg.py
29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 | |
rebin_xmet_to_vertical_track
rebin_xmet_to_vertical_track(
ds_xmet,
ds_vert,
vars=None,
k=4,
eps=1e-12,
lat_var=TRACK_LAT_VAR,
lon_var=TRACK_LON_VAR,
time_var=TIME_VAR,
height_var=HEIGHT_VAR,
along_track_dim=ALONG_TRACK_DIM,
height_dim=VERTICAL_DIM,
xmet_lat_var="latitude",
xmet_lon_var="longitude",
xmet_height_var="geometrical_height",
xmet_height_dim="height",
xmet_horizontal_grid_dim="horizontal_grid",
)
Rebins variables from an AUX_MET_1D (XMET) dataset onto the vertical curtain track of given by another dataset (e.g. ATL_EBD_2A).
This function interpolates selected variables from ds_xmet onto a EarthCARE
vertical track given in ds_vert, using quick horizontal kd-tree nearest-neighbor search with scipy.spatial.cKDTree followed
by averaging the k-nearest vertical XMET profiles using inverse distance weighting. The resulting
profiles are then interpolated in the vertical to match the height resolution of ds_vert.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds_xmet
|
Dataset | str
|
The source XMET dataset from which vertical curtain along track will be interpolated. |
required |
ds_vert
|
Dataset | str
|
The target dataset containing the vertical curtain track. |
required |
vars
|
list[str] | None
|
List of variable names from |
None
|
k
|
int
|
Number of nearest horizontal neighbors to include in the kd-tree search. Defaults to 4. |
4
|
eps
|
float
|
Numerical threshold to avoid division by zero in distance calculations during the kd-tree search. Defaults to 1e-12. |
1e-12
|
lat_var
|
str
|
Name of the latitude variable in |
TRACK_LAT_VAR
|
lon_var
|
str
|
Name of the longitude variable in |
TRACK_LON_VAR
|
time_var
|
str
|
Name of the time variable in |
TIME_VAR
|
height_var
|
str
|
Name of the height variable in |
HEIGHT_VAR
|
along_track_dim
|
str
|
Name of the along-track dimension in |
ALONG_TRACK_DIM
|
height_dim
|
str
|
Name of the vertical or height dimension in |
VERTICAL_DIM
|
xmet_lat_var
|
str
|
Name of the latitude variable in |
'latitude'
|
xmet_lon_var
|
str
|
Name of the longitude variable in |
'longitude'
|
xmet_height_var
|
str
|
Name of the height variable in |
'geometrical_height'
|
xmet_height_dim
|
str
|
Name of the vertical dimension in |
'height'
|
xmet_horizontal_grid_dim
|
str
|
Name of the horizontal grid dimension in |
'horizontal_grid'
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xr.Dataset: A new dataset containing the selected XMET variables interpolated to the grid of the
vertical curtain given in |
Raises:
| Type | Description |
|---|---|
KeyError
|
If any specified variable or coordinate name is not found in |
Source code in earthcarekit/utils/read/product/_rebin_xmet_to_vertical_track.py
19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 113 114 115 116 117 118 119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 | |
save_plot
save_plot(
fig,
filename="",
filepath=None,
ds=None,
ds_filepath=None,
pad=0.1,
dpi="figure",
orbit_and_frame=None,
utc_timestamp=None,
use_utc_creation_timestamp=False,
site_name=None,
hmax=None,
radius=None,
resolution=None,
extra=None,
transparent_outside=False,
verbose=True,
print_prefix="",
create_dirs=False,
transparent_background=False,
**kwargs
)
Save a figure as an image or vector graphic to a file and optionally format the file name in a structured way using EarthCARE metadata.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
figure
|
Figure | HasFigure
|
A figure object ( |
required |
filename
|
str
|
The base name of the file. Can be extended based on other metadata provided. Defaults to empty string. |
''
|
filepath
|
str | None
|
The path where the image is saved. Can be extended based on other metadata provided. Defaults to None. |
None
|
ds
|
Dataset | None
|
A EarthCARE dataset from which metadata will be taken. Defaults to None. |
None
|
ds_filepath
|
str | None
|
A path to a EarthCARE product from which metadata will be taken. Defaults to None. |
None
|
pad
|
float
|
Extra padding (i.e., empty space) around the image in inches. Defaults to 0.1. |
0.1
|
dpi
|
float | figure
|
The resolution in dots per inch. If 'figure', use the figure's dpi value. Defaults to None. |
'figure'
|
orbit_and_frame
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
utc_timestamp
|
TimestampLike | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
use_utc_creation_timestamp
|
bool
|
Whether the time of image creation should be included in the file name. Defaults to False. |
False
|
site_name
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
hmax
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
radius
|
int | float | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
resolution
|
str | None
|
Metadata used in the formatting of the file name. Defaults to None. |
None
|
extra
|
str | None
|
A custom string to be included in the file name. Defaults to None. |
None
|
transparent_outside
|
bool
|
Whether the area outside figures should be transparent. Defaults to False. |
False
|
verbose
|
bool
|
Whether the progress of image creation should be printed to the console. Defaults to True. |
True
|
print_prefix
|
str
|
A prefix string to all console messages. Defaults to "". |
''
|
create_dirs
|
bool
|
Whether images should be saved in a folder structure based on provided metadata. Defaults to False. |
False
|
transparent_background
|
bool
|
Whether the background inside and outside of figures should be transparent. Defaults to False. |
False
|
**kwargs
|
dict[str, Any]
|
Keyword arguments passed to wrapped function call of |
{}
|
Source code in earthcarekit/plot/save/simple_save.py
119 120 121 122 123 124 125 126 127 128 129 130 131 132 133 134 135 136 137 138 139 140 141 142 143 144 145 146 147 148 149 150 151 152 153 154 155 156 157 158 159 160 161 162 163 164 165 166 167 168 169 170 171 172 173 174 175 176 177 178 179 180 181 182 183 184 185 186 187 188 189 190 191 192 193 194 195 196 197 198 199 200 201 202 203 204 205 206 207 208 209 210 211 212 213 214 215 216 217 218 219 220 221 222 223 224 | |
search_files_by_regex
Recursively searches for files in a directory that match a given regex pattern.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root_dirpath
|
str
|
The root directory to start the search from. |
required |
regex_pattern
|
str
|
A regular expression pattern to match file names against. |
required |
Return
list[str]: A list of absolute file paths that point to files with matching names.
Raises:
| Type | Description |
|---|---|
FileNotFoundError
|
If the root directory does not exist. |
error
|
If the given pattern is not a valid regular expression. |
Source code in earthcarekit/utils/read/search.py
search_product
search_product(
root_dirpath=None,
config=None,
file_type=None,
agency=None,
latency=None,
timestamp=None,
baseline=None,
orbit_and_frame=None,
orbit_number=None,
frame_id=None,
filename=None,
start_time=None,
end_time=None,
mode="exhaustive",
)
Searches for EarthCARE product files matching given metadata filters.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
root_dirpath
|
str
|
Root directory to search. Defaults to directory given in a configuration file. |
None
|
config
|
str | ECKConfig | None
|
Path to a |
None
|
file_type
|
str | Sequence[str]
|
Product file type(s) to match. |
None
|
agency
|
str | Sequence[str]
|
Producing agency or agencies (e.g. "ESA" or "JAXA"). |
None
|
latency
|
str | Sequence[str]
|
Data latency level(s). |
None
|
timestamp
|
TimestampLike | Sequence
|
Timestamp(s) included in the product's time coverage. |
None
|
baseline
|
str | Sequence[str]
|
Baseline version(s). |
None
|
orbit_and_frame
|
str | Sequence[str]
|
Orbit and frame identifiers. |
None
|
orbit_number
|
int, str, | Sequence
|
Orbit number(s). |
None
|
frame_id
|
str | Sequence[str]
|
Frame identifier(s). |
None
|
filename
|
str | Sequence[str]
|
Specific filename(s) or regular expression patterns to match. |
None
|
start_time
|
TimestampLike
|
First timestamp included in the product's time coverage. |
None
|
end_time
|
TimestampLike
|
Last timestamp included in the product's time coverage. |
None
|
mode
|
Literal['exhaustive', 'fast']
|
Search strategy controlling completeness vs performance; the "exhaustive" mode
recursivly scans all files under the |
'exhaustive'
|
Returns:
| Name | Type | Description |
|---|---|---|
ProductDataFrame |
ProductDataFrame
|
Filtered list of matching product files as a |
Raises:
| Type | Description |
|---|---|
FileNotFoundError
|
If root directory does not exist. |
Source code in earthcarekit/utils/read/product/_search.py
252 253 254 255 256 257 258 259 260 261 262 263 264 265 266 267 268 269 270 271 272 273 274 275 276 277 278 279 280 281 282 283 284 285 286 287 288 289 290 291 292 293 294 295 296 297 298 299 300 301 302 303 304 305 306 307 308 309 310 311 312 313 314 315 316 317 318 319 320 321 322 323 324 325 326 327 328 329 330 331 332 333 334 335 336 337 338 339 340 341 342 343 344 345 346 347 348 349 350 351 352 353 354 355 356 357 358 359 360 361 362 363 364 365 366 367 368 369 370 371 372 373 374 375 376 377 378 379 380 381 382 383 384 385 386 387 388 389 390 391 392 393 394 395 396 397 398 399 400 401 402 403 404 405 406 407 408 409 410 411 412 413 414 415 416 417 418 419 420 421 422 423 424 425 426 427 428 429 430 431 432 433 434 435 436 437 438 439 440 441 442 443 444 445 446 447 448 449 450 451 452 453 454 455 456 457 458 459 460 461 462 463 464 | |
set_config
Creates or updates the default earthcarekit configuration file.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
c
|
str | ECKConfig
|
Filepath to a configuration file (.toml) or configuration object. |
required |
verbose
|
bool
|
If True, prints a message to the console. Defaults to True. |
True
|
Source code in earthcarekit/utils/config.py
set_config_maap_token
Updates the ESA MAAP access token in the default earthcarekit configuration file.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
token
|
str
|
A temporary ESA MAAP access token (to generate it visit: https://portal.maap.eo.esa.int/ini/services/auth/token/). |
required |
Source code in earthcarekit/utils/config.py
set_config_to_maap
Sets the download backend to the ESA MAAP system in the default earthcarekit configuration file.
Source code in earthcarekit/utils/config.py
set_config_to_oads
Sets the download backend to OADS in the default earthcarekit configuration file.
Source code in earthcarekit/utils/config.py
shift_cmap
Create a colormap with its center point shifted to a specified value.
This function is useful for data with asymmetric ranges (e.g., negative min and positive max) where you want the center of the colormap to align with a specific value like zero.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
cmap
|
str | Colormap | None
|
Colormap to be modified |
required |
start
|
float
|
Lower bound of the colormap range (value between 0 and |
0.0
|
midpoint
|
float
|
New center point of the colormap (value between 0 and 1). Defaults to 0.5. For data ranging from vmin to vmax where you want the center at value v, set midpoint = 1 - vmax/(vmax + abs(vmin)) |
0.5
|
stop
|
float
|
Upper bound of the colormap range (value between |
1.0
|
name
|
str
|
Name of the new colormap. Defaults to "shifted_cmap". |
'shifted_cmap'
|
Returns:
| Name | Type | Description |
|---|---|---|
Cmap |
Cmap
|
New colormap with shifted center |
Source code in earthcarekit/plot/color/colormap/shift/_shift_cmap.py
trim_to_latitude_frame_bounds
trim_to_latitude_frame_bounds(
ds,
along_track_dim=ALONG_TRACK_DIM,
lat_var=TRACK_LAT_VAR,
frame_id=None,
add_trim_index_offset_var=True,
trim_index_offset_var="trim_index_offset",
)
Trims the dataset to the region within the latitude frame bounds.
Parameters:
| Name | Type | Description | Default |
|---|---|---|---|
ds
|
Dataset
|
Input dataset to be trimmed. |
required |
along_track_dim
|
str
|
Dimension along which to trim. Defaults to ALONG_TRACK_DIM. |
ALONG_TRACK_DIM
|
lat_var
|
str
|
Name of the latitude variable. Defaults to TRACK_LAT_VAR. |
TRACK_LAT_VAR
|
frame_id
|
str | None
|
EarthCARE frame ID (single character between "A" and "H"). If given, speeds up trimming. Defaults to None. |
None
|
add_trim_index_offset_var
|
bool
|
Whether the index offset between the original and trimmed dataset is stored in the trimmed dataset (variable: "trim_index_offset"). Defaults to True. |
True
|
Returns:
| Type | Description |
|---|---|
Dataset
|
xarray.Dataset: Trimmed dataset. |